Sodium/potassium/calcium exchanger 5 (NCKX5), also known as solute carrier family 24 member 5 (SLC24A5), is a protein that in humans is encoded by the SLC24A5 gene that has a major influence on natural skin colour variation. [5] The NCKX5 protein is a member of the potassium-dependent sodium/calcium exchanger family. Sequence variation in the SLC24A5 gene, particularly a non-synonymous SNP changing the amino acid at position 111 in NCKX5 from alanine to threonine, has been associated with differences in skin pigmentation. [6]
The SLC24A5 gene's derived threonine or Ala111Thr allele (rs1426654 [7]) has been shown to be a major factor in the light skin tone of Europeans compared to Sub-Saharan Africans, and is believed to represent as much as 25–40% of the average skin tone difference between Europeans and West Africans. [5] [8] Possibly originating as long as 19,000 years ago, it has been the subject of selection in the ancestors of Europeans as recently as within the last 5,000 years, [9] and is fixed in modern European populations. [10] [11] [12] It was introduced into Khoisan people via "back-to-Africa" migration around 2,000 years ago is partly responsible for their differing skin tone to most other African populations. [13]
Gene

The SLC24A5 gene, in humans, is located on the long (q) arm of chromosome 15 on position 21.1, from base pair 46,200,461 to base pair 46,221,881. [5]
Protein
NCKX5 is 43 k Da protein that is partially localized to the trans-Golgi network in melanocytes. Removal of the NCKX5 protein disrupts melanogenesis in human and mouse melanocytes, causing a significant reduction in melanin pigment production. Site-directed mutagenesis corresponding to a non-synonymous single nucleotide polymorphism in SLC24A5 alters a residue in NCKX5 (A111T) that is important for NCKX5 sodium-calcium exchanger activity. [6]
Effect on skin color
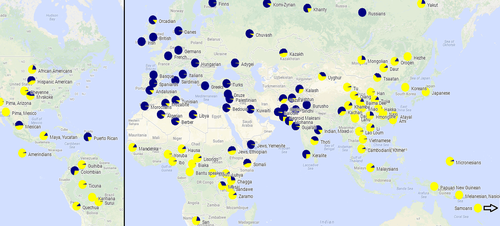
SLC24A5 appears to have played a key role in the evolution of light skin in humans of European ancestry. The gene's function in pigmentation was discovered in zebrafish as a result of the positional cloning of the gene responsible for the "golden" variety of this common pet store fish. Evidence in the International HapMap Project database of genetic variation in human populations showed that Europeans, represented by the "CEU" population, had two primary alleles differing by only one nucleotide, changing the 111th amino acid from alanine to threonine, abbreviated "A111T". [5] [14] [15]
The derived threonine allele (Ala111Thr; also known as A111T or Thr111) represented 98.7 to 100% of the alleles in European samples, while the ancestral or alanine form was found in 93 to 100% of samples of Sub-Saharan Africans, East Asians and Indigenous Americans. The variation is a SNP polymorphism rs1426654, which had been previously shown to be second among 3011 tabulated SNPs ranked as ancestry-informative markers. This single change in SLC24A5 explains between 25 and 38% of the difference in skin melanin index between peoples of sub-Saharan African and European ancestry. [5]
The SNP rs2470102 independently affects skin pigmentation variation among the South Asian population. [16]
Furthermore, the European mutation is associated with the largest region of diminished genetic variation in the CEU HapMap population, suggesting the possibility that the A111T mutation may be the subject of the single largest degree of selection in human populations of European ancestry. [5] It is hypothesized that selection for the derived allele is based on the need for sunlight to produce the essential nutrient vitamin D. In northerly latitudes, where there is less sun, greater requirement for body coverage due to colder climate, and frequently, diets poor in vitamin D, making lighter skin more suitable for survival. [17]
The earliest known sample of the threonine allele is 13,000 years old from Satsurblia Cave in Georgia. [18] The allele was widespread from Anatolia to Ukraine and Iran at the beginning of the Neolithic. [19] [20] [21]
This allele forms part of the HIrisplex DNA test system used to estimate pigmentation in forensic investigations. [22] [23]
See also
References
- ^ a b c GRCh38: Ensembl release 89: ENSG00000188467 – Ensembl, May 2017
- ^ a b c GRCm38: Ensembl release 89: ENSMUSG00000035183 – Ensembl, May 2017
- ^ "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ a b c d e f Lamason RL, Mohideen MA, Mest JR, Wong AC, Norton HL, Aros MC, et al. (December 2005). "SLC24A5, a putative cation exchanger, affects pigmentation in zebrafish and humans". Science. 310 (5755): 1782–6. Bibcode: 2005Sci...310.1782L. doi: 10.1126/science.1116238. PMID 16357253. S2CID 2245002.
- ^ a b Ginger RS, Askew SE, Ogborne RM, Wilson S, Ferdinando D, Dadd T, et al. (February 2008). "SLC24A5 encodes a trans-Golgi network protein with potassium-dependent sodium-calcium exchange activity that regulates human epidermal melanogenesis". The Journal of Biological Chemistry. 283 (9): 5486–95. doi: 10.1074/jbc.M707521200. PMID 18166528.
- ^ Reference SNP(refSNP) Cluster Report: rs1426654 **clinically associated**. Ncbi.nlm.nih.gov (2008-12-30). Retrieved on 2011-02-27.
- ^ Norton HL, Kittles RA, Parra E, McKeigue P, Mao X, Cheng K, et al. (March 2007). "Genetic evidence for the convergent evolution of light skin in Europeans and East Asians". Molecular Biology and Evolution. 24 (3): 710–22. doi: 10.1093/molbev/msl203. PMID 17182896.
- ^ Wilde S, Timpson A, Kirsanow K, Kaiser E, Kayser M, Unterländer M, et al. (April 2014). "Direct evidence for positive selection of skin, hair, and eye pigmentation in Europeans during the last 5,000 y". Proceedings of the National Academy of Sciences of the United States of America. 111 (13): 4832–7. Bibcode: 2014PNAS..111.4832W. doi: 10.1073/pnas.1316513111. PMC 3977302. PMID 24616518.
- ^ Beleza S, Santos AM, McEvoy B, Alves I, Martinho C, Cameron E, et al. (January 2013). "The timing of pigmentation lightening in Europeans". Molecular Biology and Evolution. 30 (1): 24–35. doi: 10.1093/molbev/mss207. PMC 3525146. PMID 22923467.
- ^ Soejima M, Koda Y (January 2007). "Population differences of two coding SNPs in pigmentation-related genes SLC24A5 and SLC45A2". International Journal of Legal Medicine. 121 (1): 36–9. doi: 10.1007/s00414-006-0112-z. PMID 16847698. S2CID 11192076.
- ^ Ang KC, Ngu MS, Reid KP, Teh MS, Aida ZS, Koh DX, et al. (2012). Kivisild T (ed.). "Skin color variation in Orang Asli tribes of Peninsular Malaysia". PLOS ONE. 7 (8): e42752. Bibcode: 2012PLoSO...742752A. doi: 10.1371/journal.pone.0042752. PMC 3418284. PMID 22912732.
- ^ Lin M, Siford RL, Martin AR, Nakagome S, Möller M, Hoal EG, et al. (December 2018). "Rapid evolution of a skin-lightening allele in southern African KhoeSan". Proceedings of the National Academy of Sciences of the United States of America. 115 (52): 13324–13329. Bibcode: 2018PNAS..11513324L. doi: 10.1073/pnas.1801948115. PMC 6310813. PMID 30530665.
- ^ "Key gene 'controls skin colour'". Health. BBC News. 2005-12-16. Retrieved 2010-10-23.
- ^ "Fish gene sheds light on human skin color variation". Penn State Live. Penn State University. 2005-12-16. Archived from the original on 2010-07-21. Retrieved 2010-10-23.
- ^ Prasad R, Prasad R (2016-11-20). "CCMB scientists unravel skin colour genetics of Indians". The Hindu. ISSN 0971-751X. Retrieved 2017-11-08.
- ^ Jablonski NG, Chaplin G (July 2000). "The evolution of human skin coloration" (PDF). Journal of Human Evolution. 39 (1): 57–106. doi: 10.1006/jhev.2000.0403. PMID 10896812. Archived from the original (PDF) on 2012-01-14.
- ^ Jones ER, Gonzalez-Fortes G, Connell S, Siska V, Eriksson A, Martiniano R, et al. (November 2015). "Upper Palaeolithic genomes reveal deep roots of modern Eurasians". Nature Communications. 6: 8912. Bibcode: 2015NatCo...6.8912J. doi: 10.1038/ncomms9912. PMC 4660371. PMID 26567969.
- ^ Broushaki F, Thomas MG, Link V, López S, van Dorp L, Kirsanow K, et al. (July 2016). "Early Neolithic genomes from the eastern Fertile Crescent". Science. 353 (6298): 499–503. Bibcode: 2016Sci...353..499B. doi: 10.1126/science.aaf7943. PMC 5113750. PMID 27417496.
- ^ Mathieson I, Lazaridis I, Rohland N, Mallick S, Patterson N, Roodenberg SA, Harney E, Stewardson K, Fernandes D, Novak M, Sirak K (2015-10-10). "Eight thousand years of natural selection in Europe". bioRxiv 10.1101/016477.
- ^ Jones E, Zarina G, Moiseyev V, Lightfoot E, Nigst P, Manica A, et al. (February 2017). "The Neolithic Transition in the Baltic was Not Driven by Admixture with Early European Farmers". Current Biology. 27 (4): 576–582. doi: 10.1016/j.cub.2016.12.060. PMC 5321670. PMID 28162894.
-
^ Chaitanya L, Breslin K, Zuñiga S, Wirken L, Pośpiech E, Kukla-Bartoszek M, et al. (July 2018). "The HIrisPlex-S system for eye, hair and skin colour prediction from DNA: Introduction and forensic developmental validation". Forensic Science International. Genetics. 35: 123–135.
doi:
10.1016/j.fsigen.2018.04.004.
hdl:
1805/15921.
PMID
29753263.
S2CID
21673970.
- "Forensics: New tool predicts eye, hair and skin color from a DNA sample of an unidentified individual". ScienceDaily (Press release). May 14, 2018.
- ^ Pollack A (23 February 2015). "Building a Face, and a Case, on DNA". New York Times. Retrieved 7 April 2019.
Further reading
- Grønskov K, Ek J, Sand A, Scheller R, Bygum A, Brixen K, Brondum-Nielsen K, Rosenberg T (March 2009). "Birth prevalence and mutation spectrum in danish patients with autosomal recessive albinism". Investigative Ophthalmology & Visual Science. 50 (3): 1058–64. doi: 10.1167/iovs.08-2639. PMID 19060277.
- Cook AL, Chen W, Thurber AE, Smit DJ, Smith AG, Bladen TG, et al. (February 2009). "Analysis of cultured human melanocytes based on polymorphisms within the SLC45A2/MATP, SLC24A5/NCKX5, and OCA2/P loci". The Journal of Investigative Dermatology. 129 (2): 392–405. doi: 10.1038/jid.2008.211. PMID 18650849.
- Nan H, Kraft P, Hunter DJ, Han J (August 2009). "Genetic variants in pigmentation genes, pigmentary phenotypes, and risk of skin cancer in Caucasians". International Journal of Cancer. 125 (4): 909–17. doi: 10.1002/ijc.24327. PMC 2700213. PMID 19384953.
- Meda SA, Jagannathan K, Gelernter J, Calhoun VD, Liu J, Stevens MC, et al. (November 2010). "A pilot multivariate parallel ICA study to investigate differential linkage between neural networks and genetic profiles in schizophrenia". NeuroImage. 53 (3): 1007–15. doi: 10.1016/j.neuroimage.2009.11.052. PMC 3968678. PMID 19944766.
- Stokowski RP, Pant PV, Dadd T, Fereday A, Hinds DA, Jarman C, et al. (December 2007). "A genomewide association study of skin pigmentation in a South Asian population". American Journal of Human Genetics. 81 (6): 1119–32. doi: 10.1086/522235. PMC 2276347. PMID 17999355.
- Dagle JM, Lepp NT, Cooper ME, Schaa KL, Kelsey KJ, Orr KL, et al. (April 2009). "Determination of genetic predisposition to patent ductus arteriosus in preterm infants". Pediatrics. 123 (4): 1116–23. doi: 10.1542/peds.2008-0313. PMC 2734952. PMID 19336370.
- Sulem P, Gudbjartsson DF, Stacey SN, Helgason A, Rafnar T, Magnusson KP, et al. (December 2007). "Genetic determinants of hair, eye and skin pigmentation in Europeans". Nature Genetics. 39 (12): 1443–52. doi: 10.1038/ng.2007.13. PMID 17952075. S2CID 19313549.
- Cai X, Lytton J (February 2004). "Molecular cloning of a sixth member of the K+-dependent Na+/Ca2+ exchanger gene family, NCKX6". The Journal of Biological Chemistry. 279 (7): 5867–76. doi: 10.1074/jbc.M310908200. PMID 14625281.
- Chi A, Valencia JC, Hu ZZ, Watabe H, Yamaguchi H, Mangini NJ, Huang H, Canfield VA, Cheng KC, Yang F, Abe R, Yamagishi S, Shabanowitz J, Hearing VJ, Wu C, Appella E, Hunt DF (November 2006). "Proteomic and bioinformatic characterization of the biogenesis and function of melanosomes". Journal of Proteome Research. 5 (11): 3135–44. doi: 10.1021/pr060363j. PMID 17081065.
- Soejima M, Koda Y (January 2007). "Population differences of two coding SNPs in pigmentation-related genes SLC24A5 and SLC45A2". International Journal of Legal Medicine. 121 (1): 36–9. doi: 10.1007/s00414-006-0112-z. PMID 16847698. S2CID 11192076.
- Dimisianos G, Stefanaki I, Nicolaou V, Sypsa V, Antoniou C, Poulou M, et al. (February 2009). "A study of a single variant allele (rs1426654) of the pigmentation-related gene SLC24A5 in Greek subjects". Experimental Dermatology. 18 (2): 175–7. doi: 10.1111/j.1600-0625.2008.00758.x. PMID 18637132. S2CID 22110110.
External links
- SLC24A5+protein,+human at the U.S. National Library of Medicine Medical Subject Headings (MeSH)



