| glucose transporter, type 4 | |||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||||||||||||||||||||||||||||||||||||||||||||
| Aliases | Glc_transpt_4IPR002441GLUT4Gtr4Glut-4Insulin-responsive facilitative glucose transporter | ||||||||||||||||||||||||||||||||||||||||||||||||||
| External IDs | GeneCards: [1]; OMA: - orthologs | ||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| Wikidata | |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
Glucose transporter type 4 (GLUT4), also known as solute carrier family 2, facilitated glucose transporter member 4, is a protein encoded, in humans, by the SLC2A4 gene. GLUT4 is the insulin-regulated glucose transporter found primarily in adipose tissues and striated muscle (skeletal and cardiac). The first evidence for this distinct glucose transport protein was provided by David James in 1988. [1] The gene that encodes GLUT4 was cloned [2] [3] and mapped in 1989. [4]
At the cell surface, GLUT4 permits the facilitated diffusion of circulating glucose down its concentration gradient into muscle and fat cells. Once within cells, glucose is rapidly phosphorylated by glucokinase in the liver and hexokinase in other tissues to form glucose-6-phosphate, which then enters glycolysis or is polymerized into glycogen. Glucose-6-phosphate cannot diffuse back out of cells, which also serves to maintain the concentration gradient for glucose to passively enter cells. [5]
Structure

Like all proteins, the unique amino acid arrangement in the primary sequence of GLUT4 is what allows it to transport glucose across the plasma membrane. In addition to the phenylalanine on the N-terminus, two Leucine residues and acidic motifs on the COOH-terminus are believed to play a key role in the kinetics of endocytosis and exocytosis. [7]
Other GLUT proteins
There are 14 total GLUT proteins separated into 3 classes based on sequence similarities. Class 1 consists of GLUT 1-4 and 14, class 2 contains GLUT 5, 7, 9 and 11, and class 3 has GLUT 6, 8, 10, 12 and 13.
Although there are some sequence differences between all GLUT proteins, they all have some basic structural components. For example, both the N and C termini in GLUT proteins are exposed to the cytoplasm of the cell, and they all have 12 transmembrane segments. [8]
Tissue distribution
Skeletal muscle
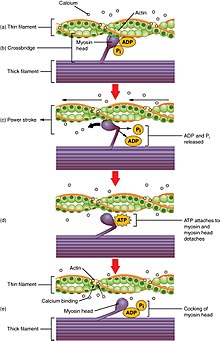
In striated skeletal muscle cells, GLUT4 concentration in the plasma membrane can increase as a result of either exercise or muscle contraction.
During exercise, the body needs to convert glucose to ATP to be used as energy. As G-6-P concentrations decrease, hexokinase becomes less inhibited, and the glycolytic and oxidative pathways that make ATP are able to proceed. This also means that muscle cells are able to take in more glucose as its intracellular concentrations decrease. In order to increase glucose levels in the cell, GLUT4 is the primary transporter used in this facilitated diffusion. [10]
Although muscle contractions function in a similar way and also induce the translocation of GLUT4 into the plasma membrane, the two skeletal muscle processes obtain different forms of intracellular GLUT4. The GLUT4 carrier vesicles are either transferrin positive or negative, and are recruited by different stimuli. Transferrin-positive GLUT4 vesicles are utilized during muscle contraction while the transferrin-negative vesicles are activated by insulin stimulation as well as by exercise. [11] [12]
Cardiac muscle
Cardiac muscle is slightly different from skeletal muscle. At rest, they prefer to utilize fatty acids as their main energy source. As activity increases and it begins to pump faster, the cardiac muscles begin to oxidize glucose at a higher rate. [13]
An analysis of mRNA levels of GLUT1 and GLUT4 in cardiac muscles show that GLUT1 plays a larger role in cardiac muscles than it does in skeletal muscles. [14] GLUT4, however, is still believed to be the primary transporter for glucose. [15]
Much like in other tissues, GLUT4 also responds to insulin signaling, and is transported into the plasma membrane to facilitate the diffusion of glucose into the cell. [16] [17]
Adipose tissue
Adipose tissue, commonly known as fat, [18] is a depository for energy in order to conserve metabolic homeostasis. As the body takes in energy in the form of glucose, some is expended, and the rest is stored as glycogen (primarily in the liver, muscle cells), or as triglyceride in adipose tissue. [19]
An imbalance in glucose intake and energy expenditure has been shown to lead to both adipose cell hypertrophy and hyperplasia, which lead to obesity. [20] In addition, mutations in GLUT4 genes in adipocytes can also lead to increased GLUT4 expression in adipose cells, which allows for increased glucose uptake and therefore more fat stored. If GLUT4 is over-expressed, it can actually alter nutrient distribution and send excess glucose into adipose tissue, leading to increased adipose tissue mass. [20]
Regulation
Insulin
Insulin is released from the pancreas and into the bloodstream in response to increased glucose concentration in the blood. [21] Insulin is stored in beta cells in the pancreas. When glucose in the blood binds to glucose receptors on the beta cell membrane, a signal cascade is initiated inside the cell that results in insulin stored in vesicles in these cells being released into the blood stream. [22] Increased insulin levels cause the uptake of glucose into the cells. GLUT4 is stored in the cell in transport vesicles, and is quickly incorporated into the plasma membrane of the cell when insulin binds to membrane receptors. [19]
Under conditions of low insulin, most GLUT4 is sequestered in intracellular vesicles in muscle and fat cells. As the vesicles fuse with the plasma membrane, GLUT4 transporters are inserted and become available for transporting glucose, and glucose absorption increases. [23] The genetically engineered muscle insulin receptor knock‐out (MIRKO) mouse was designed to be insensitive to glucose uptake caused by insulin, meaning that GLUT4 is absent. Mice with diabetes or fasting hyperglycemia, however, were found to be immune to the negative effects of the insensitivity. [24]

The mechanism for GLUT4 is an example of a cascade effect, where binding of a ligand to a membrane receptor amplifies the signal and causes a cellular response. In this case, insulin binds to the insulin receptor in its dimeric form and activates the receptor's tyrosine-kinase domain. The receptor then recruits Insulin Receptor Substrate, or IRS-1, which binds the enzyme PI-3 kinase. PI-3 kinase converts the membrane lipid PIP2 to PIP3. PIP3 is specifically recognized by PKB ( protein kinase B) and by PDK1, which can phosphorylate and activate PKB. Once phosphorylated, PKB is in its active form and phosphorylates TBC1D4, which inhibits the GTPase-activating domain associated with TBC1D4, allowing for Rab protein to change from its GDP to GTP bound state. Inhibition of the GTPase-activating domain leaves proteins next in the cascade in their active form, and stimulates GLUT4 to be expressed on the plasma membrane. [25]
RAC1 is a GTPase also activated by insulin. Rac1 stimulates reorganization of the cortical Actin cytoskeleton [26] which allows for the GLUT4 vesicles to be inserted into the plasma membrane. [27] [28] A RAC1 Knockout mouse has reduced glucose uptake in muscle tissue. [28]
Knockout mice that are heterozygous for GLUT4 develop insulin resistance in their muscles as well as diabetes. [29]
Muscle contraction
Muscle contraction stimulates muscle cells to translocate GLUT4 receptors to their surfaces. This is especially true in cardiac muscle, where continuous contraction increases the rate of GLUT4 translocation; but is observed to a lesser extent in increased skeletal muscle contraction. [30] In skeletal muscle, muscle contractions increase GLUT4 translocation severalfold, [31] and this is likely regulated by RAC1 [32] [33] and AMP-activated protein kinase. [34]
Muscle stretching
Muscle stretching also stimulates GLUT4 translocation and glucose uptake in rodent muscle via RAC1. [35]
Interactions
GLUT4 has been shown to interact with death-associated protein 6, also known as Daxx. Daxx, which is used to regulate apoptosis, has been shown to associate with GLUT4 in the cytoplasm. UBX-domains, such as the one found in GLUT4, have been shown to associate with apoptotic signaling. [6] So this interaction aids in the translocation of Daxx within the cell. [36]
In addition, recent reports demonstrated the presence of GLUT4 gene in central nervous system such as the hippocampus. Moreover, impairment in insulin-stimulated trafficking of GLUT4 in the hippocampus result in decreased metabolic activities and plasticity of hippocampal neurons, which leads to depressive like behaviour and cognitive dysfunction. [37] [38] [39]
Interactive pathway map
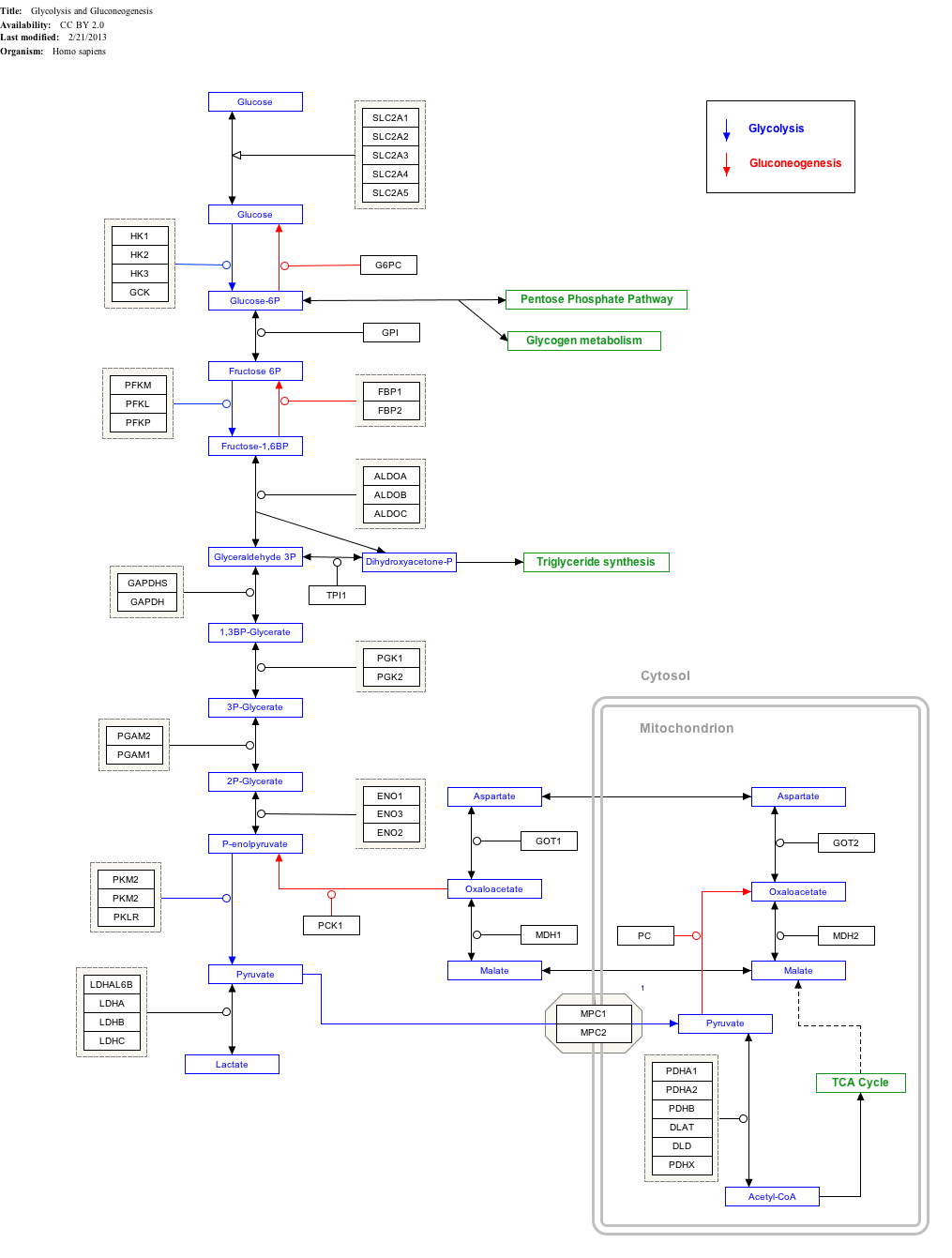
Click on genes, proteins and metabolites below to link to respective articles. [§ 1]
- ^ The interactive pathway map can be edited at WikiPathways: "GlycolysisGluconeogenesis_WP534".
References
- ^ James DE, Brown R, Navarro J, Pilch PF (May 1988). "Insulin-regulatable tissues express a unique insulin-sensitive glucose transport protein". Nature. 333 (6169): 183–5. Bibcode: 1988Natur.333..183J. doi: 10.1038/333183a0. PMID 3285221. S2CID 4237493.
- ^ James DE, Strube M, Mueckler M (March 1989). "Molecular cloning and characterization of an insulin-regulatable glucose transporter". Nature. 338 (6210): 83–7. Bibcode: 1989Natur.338...83J. doi: 10.1038/338083a0. PMID 2645527. S2CID 4285627.
- ^ Birnbaum MJ (April 1989). "Identification of a novel gene encoding an insulin-responsive glucose transporter protein". Cell. 57 (2): 305–15. doi: 10.1016/0092-8674(89)90968-9. PMID 2649253. S2CID 20359706.
- ^ Bell GI, Murray JC, Nakamura Y, Kayano T, Eddy RL, Fan YS, Byers MG, Shows TB (August 1989). "Polymorphic human insulin-responsive glucose-transporter gene on chromosome 17p13". Diabetes. 38 (8): 1072–5. doi: 10.2337/diabetes.38.8.1072. PMID 2568955.
- ^ Watson RT, Kanzaki M, Pessin JE (April 2004). "Regulated membrane trafficking of the insulin-responsive glucose transporter 4 in adipocytes". Endocrine Reviews. 25 (2): 177–204. doi: 10.1210/er.2003-0011. PMID 15082519.
- ^ a b Buchberger A, Howard MJ, Proctor M, Bycroft M (March 2001). "The UBX domain: a widespread ubiquitin-like module". Journal of Molecular Biology. 307 (1): 17–24. doi: 10.1006/jmbi.2000.4462. PMID 11243799.
- ^ Huang S, Czech MP (April 2007). "The GLUT4 glucose transporter". Cell Metabolism. 5 (4): 237–52. doi: 10.1016/j.cmet.2007.03.006. PMID 17403369.
- ^ Mueckler M, Thorens B (2013). "The SLC2 (GLUT) family of membrane transporters". Molecular Aspects of Medicine. 34 (2–3): 121–38. doi: 10.1016/j.mam.2012.07.001. PMC 4104978. PMID 23506862.
- ^ Lodish H, Berk A, Zipursky SL, Matsudaira P, Baltimore D, Darnell J (2000). "16.1: Oxidation of Glucose and Fatty Acids to CO2". Molecular Cell Biology (4th ed.). New York: W. H. Freeman. ISBN 978-0-7167-3706-3.
- ^ Richter EA, Hargreaves M (July 2013). "Exercise, GLUT4, and skeletal muscle glucose uptake". Physiological Reviews. 93 (3): 993–1017. doi: 10.1152/physrev.00038.2012. PMID 23899560.
- ^ Ploug T, van Deurs B, Ai H, Cushman SW, Ralston E (September 1998). "Analysis of GLUT4 distribution in whole skeletal muscle fibers: identification of distinct storage compartments that are recruited by insulin and muscle contractions". The Journal of Cell Biology. 142 (6): 1429–46. doi: 10.1083/jcb.142.6.1429. PMC 2141761. PMID 9744875.
- ^ Lauritzen HP (April 2013). "Insulin- and contraction-induced glucose transporter 4 traffic in muscle: insights from a novel imaging approach". Exercise and Sport Sciences Reviews. 41 (2): 77–86. doi: 10.1097/JES.0b013e318275574c. PMC 3602324. PMID 23072821.
- ^ Morgan HE, Henderson MJ, Regen DM, Park CR (September 1959). "Regulation of glucose uptake in heart muscle from normal and alloxan-diabetic rats: the effects of insulin, growth hormone, cortisone, and anoxia". Annals of the New York Academy of Sciences. 82 (2): 387–402. Bibcode: 1959NYASA..82..387M. doi: 10.1111/j.1749-6632.1959.tb44920.x. PMID 14424107. S2CID 32458568.
- ^ Laybutt DR, Thompson AL, Cooney GJ, Kraegen EW (September 1997). "Selective chronic regulation of GLUT1 and GLUT4 content by insulin, glucose, and lipid in rat cardiac muscle in vivo". The American Journal of Physiology. 273 (3 Pt 2): H1309–16. doi: 10.1152/ajpheart.1997.273.3.H1309. PMID 9321820.
- ^ Rett K, Wicklmayr M, Dietze GJ, Häring HU (January 1996). "Insulin-induced glucose transporter (GLUT1 and GLUT4) translocation in cardiac muscle tissue is mimicked by bradykinin". Diabetes. 45 Suppl 1 (Supplement 1): S66–9. doi: 10.2337/diab.45.1.S66. PMID 8529803. S2CID 7766813.
- ^ Slot JW, Geuze HJ, Gigengack S, James DE, Lienhard GE (September 1991). "Translocation of the glucose transporter GLUT4 in cardiac myocytes of the rat". Proceedings of the National Academy of Sciences of the United States of America. 88 (17): 7815–9. Bibcode: 1991PNAS...88.7815S. doi: 10.1073/pnas.88.17.7815. PMC 52394. PMID 1881917.
- ^ Luiken JJ, Glatz JF, Neumann D (August 2015). "Cardiac contraction-induced GLUT4 translocation requires dual signaling input" (PDF). Trends Endocrinol Metab. 26 (8): 404–10. doi: 10.1016/j.tem.2015.06.002. PMID 26138758. S2CID 171571.
- ^ "Adipose tissue". ScienceDaily. Retrieved 2017-05-24.
- ^ a b Favaretto F, Milan G, Collin GB, Marshall JD, Stasi F, Maffei P, Vettor R, Naggert JK (2014-10-09). "GLUT4 defects in adipose tissue are early signs of metabolic alterations in Alms1GT/GT, a mouse model for obesity and insulin resistance". PLOS ONE. 9 (10): e109540. Bibcode: 2014PLoSO...9j9540F. doi: 10.1371/journal.pone.0109540. PMC 4192353. PMID 25299671.
- ^ a b Shepherd PR, Gnudi L, Tozzo E, Yang H, Leach F, Kahn BB (October 1993). "Adipose cell hyperplasia and enhanced glucose disposal in transgenic mice overexpressing GLUT4 selectively in adipose tissue". The Journal of Biological Chemistry. 268 (30): 22243–6. doi: 10.1016/S0021-9258(18)41516-5. PMID 8226728.
- ^ "Insulin Synthesis and Secretion". www.vivo.colostate.edu. Retrieved 2017-05-24.
- ^ Fu, Zhuo (2013). "Regulation of Insulin Synthesis and Secretion and Pancreatic Beta-Cell Dysfunction in Diabetes". Curr Diabetes Rev. 9 (1): 25–53. doi: 10.2174/1573399811309010025. PMC 3934755. PMID 22974359.
- ^ Cushman SW, Wardzala LJ (May 1980). "Potential mechanism of insulin action on glucose transport in the isolated rat adipose cell. Apparent translocation of intracellular transport systems to the plasma membrane" (PDF). The Journal of Biological Chemistry. 255 (10): 4758–62. doi: 10.1016/S0021-9258(19)85561-8. PMID 6989818.
- ^ Sonksen P, Sonksen J (July 2000). "Insulin: understanding its action in health and disease". British Journal of Anaesthesia. 85 (1): 69–79. doi: 10.1093/bja/85.1.69. PMID 10927996.
- ^ Leto, Dara; Saltiel, Alan R. (May 2012). "Regulation of glucose transport by insulin: traffic control of GLUT4". Nature Reviews Molecular Cell Biology. 13 (6): 383–396. doi: 10.1038/nrm3351. ISSN 1471-0072. PMID 22617471. S2CID 39756994.
- ^ JeBailey L, Wanono O, Niu W, Roessler J, Rudich A, Klip A (February 2007). "Ceramide- and oxidant-induced insulin resistance involve loss of insulin-dependent Rac-activation and actin remodeling in muscle cells". Diabetes. 56 (2): 394–403. doi: 10.2337/db06-0823. PMID 17259384.
- ^ Sylow L, Kleinert M, Pehmøller C, Prats C, Chiu TT, Klip A, Richter EA, Jensen TE (February 2014). "Akt and Rac1 signaling are jointly required for insulin-stimulated glucose uptake in skeletal muscle and downregulated in insulin resistance". Cellular Signalling. 26 (2): 323–31. doi: 10.1016/j.cellsig.2013.11.007. PMID 24216610.
- ^ a b Sylow L, Jensen TE, Kleinert M, Højlund K, Kiens B, Wojtaszewski J, Prats C, Schjerling P, Richter EA (June 2013). "Rac1 signaling is required for insulin-stimulated glucose uptake and is dysregulated in insulin-resistant murine and human skeletal muscle". Diabetes. 62 (6): 1865–75. doi: 10.2337/db12-1148. PMC 3661612. PMID 23423567.
- ^ Stenbit AE, Tsao TS, Li J, Burcelin R, Geenen DL, Factor SM, Houseknecht K, Katz EB, Charron MJ (October 1997). "GLUT4 heterozygous knockout mice develop muscle insulin resistance and diabetes". Nature Medicine. 3 (10): 1096–101. doi: 10.1038/nm1097-1096. PMID 9334720. S2CID 8643507.
- ^ Lund S, Holman GD, Schmitz O, Pedersen O (June 1995). "Contraction stimulates translocation of glucose transporter GLUT4 in skeletal muscle through a mechanism distinct from that of insulin". Proceedings of the National Academy of Sciences of the United States of America. 92 (13): 5817–21. Bibcode: 1995PNAS...92.5817L. doi: 10.1073/pnas.92.13.5817. PMC 41592. PMID 7597034.
- ^ Jensen TE, Sylow L, Rose AJ, Madsen AB, Angin Y, Maarbjerg SJ, Richter EA (October 2014). "Contraction-stimulated glucose transport in muscle is controlled by AMPK and mechanical stress but not sarcoplasmatic reticulum Ca(2+) release". Molecular Metabolism. 3 (7): 742–53. doi: 10.1016/j.molmet.2014.07.005. PMC 4209358. PMID 25353002.
- ^ Sylow L, Møller LL, Kleinert M, Richter EA, Jensen TE (December 2014). "Rac1--a novel regulator of contraction-stimulated glucose uptake in skeletal muscle". Experimental Physiology. 99 (12): 1574–80. doi: 10.1113/expphysiol.2014.079194. PMID 25239922.
- ^ Sylow L, Jensen TE, Kleinert M, Mouatt JR, Maarbjerg SJ, Jeppesen J, Prats C, Chiu TT, Boguslavsky S, Klip A, Schjerling P, Richter EA (April 2013). "Rac1 is a novel regulator of contraction-stimulated glucose uptake in skeletal muscle". Diabetes. 62 (4): 1139–51. doi: 10.2337/db12-0491. PMC 3609592. PMID 23274900.
- ^ Mu J, Brozinick JT, Valladares O, Bucan M, Birnbaum MJ (May 2001). "A role for AMP-activated protein kinase in contraction- and hypoxia-regulated glucose transport in skeletal muscle". Molecular Cell. 7 (5): 1085–94. doi: 10.1016/s1097-2765(01)00251-9. PMID 11389854.
- ^ Sylow L, Møller LL, Kleinert M, Richter EA, Jensen TE (February 2015). "Stretch-stimulated glucose transport in skeletal muscle is regulated by Rac1". The Journal of Physiology. 593 (3): 645–56. doi: 10.1113/jphysiol.2014.284281. PMC 4324711. PMID 25416624.
- ^ Lalioti VS, Vergarajauregui S, Pulido D, Sandoval IV (May 2002). "The insulin-sensitive glucose transporter, GLUT4, interacts physically with Daxx. Two proteins with capacity to bind Ubc9 and conjugated to SUMO1". The Journal of Biological Chemistry. 277 (22): 19783–91. doi: 10.1074/jbc.M110294200. PMID 11842083.
- ^ Patel SS, Udayabanu M (March 2014). "Urtica dioica extract attenuates depressive like behavior and associative memory dysfunction in dexamethasone induced diabetic mice". Metabolic Brain Disease. 29 (1): 121–30. doi: 10.1007/s11011-014-9480-0. PMID 24435938. S2CID 10955351.
- ^ Piroli GG, Grillo CA, Reznikov LR, Adams S, McEwen BS, Charron MJ, Reagan LP (2007). "Corticosterone impairs insulin-stimulated translocation of GLUT4 in the rat hippocampus". Neuroendocrinology. 85 (2): 71–80. doi: 10.1159/000101694. PMID 17426391. S2CID 38081413.
- ^ Huang CC, Lee CC, Hsu KS (2010). "The role of insulin receptor signaling in synaptic plasticity and cognitive function". Chang Gung Medical Journal. 33 (2): 115–25. PMID 20438663.
External links
- GLUT4+Protein at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
- USCD—Nature molecule pages: The signaling pathway", "GLUT4"; contains a high-resolution network map. Accessed 25 December 2009.