| Retron msr RNA | |
|---|---|
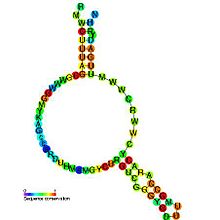 Predicted
secondary structure and
sequence conservation of msr | |
| Identifiers | |
| Symbol | msr |
| Rfam | RF00170 |
| Other data | |
| RNA type | Gene |
| Domain(s) | Bacteria |
| SO | SO:0000233 |
| PDB structures | PDBe |
A retron is a distinct DNA sequence found in the genome of many bacteria species that codes for reverse transcriptase and a unique single-stranded DNA/RNA hybrid called multicopy single-stranded DNA (msDNA). Retron msr RNA is the non-coding RNA produced by retron elements and is the immediate precursor to the synthesis of msDNA. The retron msr RNA folds into a characteristic secondary structure that contains a conserved guanosine residue at the end of a stem loop. Synthesis of DNA by the retron-encoded reverse transcriptase (RT) results in a DNA/RNA chimera which is composed of small single-stranded DNA linked to small single-stranded RNA. The RNA strand is joined to the 5′ end of the DNA chain via a 2′–5′ phosphodiester linkage that occurs from the 2′ position of the conserved internal guanosine residue.
Sequence and structure

Retron elements are about 2 kb long. They contain a single operon controlling the synthesis of an RNA transcript carrying three loci, msr, msd, and ret, that are involved in msDNA synthesis. The DNA portion of msDNA is encoded by the msd gene, the RNA portion is encoded by the msr gene, while the product of the ret gene is a reverse transcriptase similar to the RTs produced by retroviruses and other types of retroelements. [1] Like other reverse transcriptases, the retron RT contains seven regions of conserved amino acids (labeled 1–7 in the figure), including a highly conserved tyr- ala- asp- asp (YADD) sequence associated with the catalytic core. The ret gene product is responsible for processing the msd/msr portion of the RNA transcript into msDNA.
Classification and occurrence
For many years after their discovery in animal viruses, reverse transcriptases were believed to be absent from prokaryotes. Currently, however, RT-encoding elements, i.e. retroelements, have been found in a wide variety of different bacteria:
- Retrons were the first family of retroelement discovered in bacteria; the other two families of known bacterial retroelements are:
- group II introns: Group II introns are the best characterized bacterial retroelement and the only type known to exhibit autonomous mobility; they consist of an RT encoded within a catalytic, self-splicing RNA structure. Group II intron mobility is mediated by a ribonucleoprotein comprising an intron lariat bound to two intron-coded proteins. [2]
- diversity-generating retroelements (DGRs). [3] The DGRs are not mobile, but function to diversify DNA sequences. [2] For example, DGRs mediate the switch between pathogenic and free-living phases of Bordetella. [4]
Function
Since retrons are not mobile, their appearance in diverse bacterial species is not a " selfish DNA" phenomenon. Rather, bacterial retrons confer some protection from phage infection to bacterial hosts. Several retrons are located in DNA regions next to certain protein effector-coding genes. When their expression is activated, most of these effectors and their associated retrons function together to block phage infection. [5] [6]
Retrons are being developed into genome-editing tools. [7]
References
- ^ Lampson BC, Inouye M, Inouye S (2005). "Retrons, msDNA, and the bacterial genome" (PDF). Cytogenet Genome Res. 110 (1–4): 491–499. doi: 10.1159/000084982. PMID 16093702. S2CID 24854188.
- ^ a b Medhekar B, Mille JF (2007). "Diversity-Generating Retroelements". Current Opinion in Microbiology. 10 (4): 388–395. doi: 10.1016/j.mib.2007.06.004. PMC 2703298. PMID 17703991.
- ^ Simon DM, Zimmerly S (2008). "A diversity of uncharacterized reverse transcriptases in bacteria". Nucleic Acids Res. 36 (22): 7219–7229. CiteSeerX 10.1.1.358.8390. doi: 10.1093/nar/gkn867. PMC 2602772. PMID 19004871.
- ^ Liu M, Gingery M, Doulatov SR, Liu Y, Hodes A, Baker S, Davis P, Simmonds M, Churcher C, Mungall K, Quail MA, Preston A, Harvill ET, Maskell DJ, Eiserling FA, Parkhill J, Miller JF (2004). "Genomic and Genetic Analysis of Bordetella Bacteriophages Encoding Reverse Transcriptase-Mediated Tropism-Switching Cassettes". J. Bacteriol. 186 (5): 1503–1517. doi: 10.1128/JB.186.5.1503-1517.2004. PMC 344406. PMID 14973019.
- ^ Bobonis, Jacob; Mitosch, Karin; Mateus, André; Karcher, Nicolai; Kritikos, George; Selkrig, Joel; Zietek, Matylda; Monzon, Vivian; Pfalz, Birgit; Garcia-Santamarina, Sarela; Galardini, Marco; Sueki, Anna; Kobayashi, Callie; Stein, Frank; Bateman, Alex (2022-09-01). "Bacterial retrons encode phage-defending tripartite toxin–antitoxin systems". Nature. 609 (7925): 144–150. doi: 10.1038/s41586-022-05091-4. ISSN 0028-0836. PMID 35850148. S2CID 250643138.
- ^ Millman A, Bernheim A, Stokar-Avihail A, Fedorenko T, Voichek M, Leavitt A, Oppenheimer-Shaanan Y, Sorek R (2020). "Bacterial Retrons Function In Anti-Phage Defense". Cell. 183 (6): 1551–1561. doi: 10.1016/j.cell.2020.09.065. PMID 33157039.
- ^ Simon AJ, Ellington AD, Finkelstein IJ (2019). "Retrons and their applications in genome engineering". Nucleic Acids Research. 47 (21): 11007–11019. doi: 10.1093/nar/gkz865. PMC 6868368. PMID 31598685.
External links
- Page for Retron msr RNA at Rfam
- Mystery molecule in bacteria is revealed to be a guard, on: EurekAlert!, 5 Nov 2020. Source: WEIZMANN INSTITUTE OF SCIENCE