| Cyclohexanone monooxygenase | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||
| EC no. | 1.14.13.22 | ||||||||
| CAS no. | 52037-90-8 | ||||||||
| Databases | |||||||||
| IntEnz | IntEnz view | ||||||||
| BRENDA | BRENDA entry | ||||||||
| ExPASy | NiceZyme view | ||||||||
| KEGG | KEGG entry | ||||||||
| MetaCyc | metabolic pathway | ||||||||
| PRIAM | profile | ||||||||
| PDB structures | RCSB PDB PDBe PDBsum | ||||||||
| |||||||||
Cyclohexanone monooxygenase ( EC 1.14.13.22, cyclohexanone 1,2-monooxygenase, cyclohexanone oxygenase, cyclohexanone:NADPH:oxygen oxidoreductase (6-hydroxylating, 1,2-lactonizing)) is an enzyme with systematic name cyclohexanone,NADPH:oxygen oxidoreductase (lactone-forming). [1] [2] [3] [4] [5] [6] This enzyme catalyses the following chemical reaction
- cyclohexanone + NADPH + H+ + O2 hexano-6-lactone + NADP+ + H2O
This enzyme contains 540 residues organized into a single subunit. Cyclohexanone monooxygenase is one of the most prominent Baeyer-Villiger monooxygenases (BVMOs) and has a low substrate specificity, allowing it to catalyze a number of reactions; given the variety of substrates, cyclohexanone monooxygenase is a useful enzyme for industrial applications.
Enzyme mechanism
Cyclohexanone monooxygenase (CHMO) uses NADPH and O2 as cosubstrates and FAD as a cofactor to insert an oxygen atom into the substrate. The process involves the formation of a falvin-peroxide and Criegee intermediate. [7]
CHMO is a member of the Baeyer-Villiger monooxygenase (BVMO) family and flavin-containing monooxygenases (FMO) superfamily. [7]
Cyclohexanone undergoes the following process, similar to the Baeyer-Villiger reactions, to be converted into hexano-6-lactone using CHMO. [8]
- NADPH attaches to the active site of CHMO and transfers a hydride resulting in FADH- and NADP+.
- A one-electron transfer from FADH- to O2 results in a superoxide radical and FAD semiquinone.
- Recombination of the radical pair results in the C4a-peroxyflavin intermediate.
- The C4a-peroxyflavin intermediate functions as a nucleophile and attacks the cyclohexanone substrate to form the Criegee intermediate.
- The intermediate undergoes a rearrangement to produce the hexano-6-lactone.
- CHMO releases H2O and NADP+ and regenerates FADH.
CHMO can also oxygenate cyclic ketones, aromatic aldehydes, and heteroatom-containing compounds. [9]
Enzyme structure
Using CHMO isolated from Rhodococcus sp. Strain HI-31 and complexed with FAD and NADP+, two crystal structures were obtained showing CHMO in the open and closed conformations. [7] Structurally, CHMO is stable and contains 540 residues organized into a single subunit.
CHMO contains binding domains for NADP+ and FAD, which are connected by two unstructured loops. The NADP binding domain consists of the segments 152-208 and 335-380 with a helical domain constructed between residues 224-332. The helical domain shifts between the two dinucleotide ( NADP+ and FAD) binding domains and helps form the substrate binding pocket. The FAD binding domain consists of the first 140 N-terminal residues as well as residues 387-540 from the C-terminus. [7]

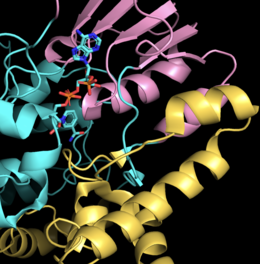
The substrate binding pocket is well defined in the closed conformation and consists of the residues 145−146, 248, 279, 329, 434−435, 437, 492, and 507; FAD and NADP+ also contribute to the shape of the binding pocket. [7]
The key distinction between the open form, CHMOopen, and the closed form, CHMOclosed, lies in the conformation of residues 487-504, which form a loop. In the closed confirmation, the loop folds upon itself, internalizing the center portion of the loop. However, in the open conformation, the loop is not visible. It is predicted that this results from the loop adopting a solvent-exposed conformation. [7]

Biological function
CHMO is a bacterial flavoenzyme whose main function in the cell is to catalyze the conversion of cyclohexanone, a cyclic ketone, into ε- caprolactone, a key step in the pathway for the biodedgredation of cyclohexanol. [10] However, given the lack of specificity for CHMO, it can be used generally to form lactones from a number of four to six-membered cyclic ketones, which can then be hydrolyzed into aliphatic acids. [10] Moreover, CHMO has the ability to oxygenate aromatic aldehydes and heteroatom-containing compounds – such as trivalent phosphorus and boronic acids– as well, making it a candidate for industrial use. [10]
Industrial relevance
Utilizing its affinity for multiple substrates and given that the mechanism is one of the most well studied Baeyer-Villiger Monooxygenases (BVMOs) with high regio-, chemo- and enantioselectivity, CHMO has been identified as a useful industrial molecule. [11] [7] Strain-specific primers derived from the CMHO gene have already been used to developed and optimized to both quantify and monitor levels of Lysobacter antibioticus, a potential biological disease control for crops, in agricultural soils by PCR and real-time qPCR. [12] With regard to the healthcare industry, CHMO mutants are a candidate for the efficient extraceullular enzymatic synthesis of (S)- omeprazole– a drug for gastroesophageal refux– when expressed by Pichia pastoris, a methylotrophic yeast. [13] Additionally, CHMO has demonstrated its ability to form chiral synthons making CHMO a potential target for more cost-effective drug synthesis, specifically with regard to enantioselective lactones. [10]
References
- ^ Donoghue NA, Norris DB, Trudgill PW (March 1976). "The purification and properties of cyclohexanone oxygenase from Nocardia globerula CL1 and Acinetobacter NCIB 9871". European Journal of Biochemistry. 63 (1): 175–92. doi: 10.1111/j.1432-1033.1976.tb10220.x. PMID 1261545.
- ^ Sheng D, Ballou DP, Massey V (September 2001). "Mechanistic studies of cyclohexanone monooxygenase: chemical properties of intermediates involved in catalysis". Biochemistry. 40 (37): 11156–67. doi: 10.1021/bi011153h. PMID 11551214.
- ^ Stewart, J.D. (1998). "Cyclohexanone monooxygenase: a useful reagent for asymmetric Baeyer-Villiger reactions". Curr. Org. Chem. 2 (3): 195–216. doi: 10.2174/1385272802666220128191443.
- ^ Kayser M, Mihovilovic M, Mrstik M, Martinez C, Stewart J (1999). "Asymmetric oxidations at sulfur catalyzed by engineered strains that overexpress cyclohexanone monooxygenase". New Journal of Chemistry. 23 (8): 827–832. doi: 10.1039/a902283j.
- ^ Ottolina G, Bianchi S, Belloni B, Carrea G, Danieli B (1999). "First asymmetric oxidation of tertiary amines by cyclohexanone monooxygenase". Tetrahedron Lett. 40 (48): 8483–8486. doi: 10.1016/s0040-4039(99)01780-3.
- ^ Colonna S, Gaggero N, Carrea G, Ottolina G, Pasta P, Zambianchi F (2002). "First asymmetric epoxidation catalysed by cyclohexanone monooxygenase". Tetrahedron Lett. 43 (10): 1797–1799. doi: 10.1016/s0040-4039(02)00029-1.
- ^ a b c d e f g Mirza, I. Ahmad; Yachnin, Brahm J.; Wang, Shaozhao; Grosse, Stephan; Bergeron, Hélène; Imura, Akihiro; Iwaki, Hiroaki; Hasegawa, Yoshie; Lau, Peter C. K.; Berghuis, Albert M. (2009-07-01). "Crystal Structures of Cyclohexanone Monooxygenase Reveal Complex Domain Movements and a Sliding Cofactor". Journal of the American Chemical Society. 131 (25): 8848–8854. doi: 10.1021/ja9010578. ISSN 0002-7863. PMID 19385644. S2CID 207138937.
- ^ Fordwour, Osei Boakye; Wolthers, Kirsten R. (2018-12-01). "Active site arginine controls the stereochemistry of hydride transfer in cyclohexanone monooxygenase". Archives of Biochemistry and Biophysics. 659: 47–56. doi: 10.1016/j.abb.2018.09.025. ISSN 0003-9861. PMID 30287236. S2CID 52919877.
- ^ Sheng, D.; Ballou, D. P.; Massey, V. (2001-09-18). "Mechanistic studies of cyclohexanone monooxygenase: chemical properties of intermediates involved in catalysis". Biochemistry. 40 (37): 11156–11167. doi: 10.1021/bi011153h. ISSN 0006-2960. PMID 11551214.
- ^ a b c d Sheng, Dawei; Ballou, David P.; Massey, Vincent (2001-09-01). "Mechanistic Studies of Cyclohexanone Monooxygenase: Chemical Properties of Intermediates Involved in Catalysis". Biochemistry. 40 (37): 11156–11167. doi: 10.1021/bi011153h. ISSN 0006-2960. PMID 11551214.
- ^ Beek, Hugo L. van; Gonzalo, Gonzalo de; Fraaije, Marco W. (2012-03-05). "Blending Baeyer–Villiger monooxygenases: using a robust BVMO as a scaffold for creating chimeric enzymes with novel catalytic properties". Chemical Communications. 48 (27): 3288–3290. doi: 10.1039/C2CC17656D. ISSN 1364-548X. PMID 22286124.
- ^ Lina, Fu; Ting, Wang; Lanfang, Wei; Jun, Yang; Qi, Liu; Yating, Wang; Xing, Wang; Guanghai, Ji (2018). "Specific detection of Lysobacter antibioticus strains in agricultural soil using PCR and real-time PCR". FEMS Microbiology Letters. 365 (20). doi: 10.1093/femsle/fny219. PMID 30202922.
- ^ Li, Ya-Jing; Zheng, Yu-Cong; Geng, Qiang; Liu, Feng; Zhang, Zhi-Jun; Xu, Jian-He; Yu, Hui-Lei (2021-08-27). "Secretory expression of cyclohexanone monooxygenase by methylotrophic yeast for efficient omeprazole sulfide bio-oxidation". Bioresources and Bioprocessing. 8 (1): 81. doi: 10.1186/s40643-021-00430-1. ISSN 2197-4365. PMC 10992682. S2CID 237330516.
External links
- Cyclohexanone+monooxygenase at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
