| Rhodobacter sphaeroides | |
|---|---|
| |
| Rhodobacter sphaeroides | |
| Scientific classification | |
| Domain: | |
| Phylum: | |
| Class: | |
| Order: | |
| Family: | |
| Genus: | |
| Species: | R. sphaeroides
|
| Binomial name | |
| Rhodobacter sphaeroides (van Niel, 1944) Imhoff et al., 1984
| |
Rhodobacter sphaeroides is a kind of purple bacterium; a group of bacteria that can obtain energy through photosynthesis. Its best growth conditions are anaerobic phototrophy ( photoheterotrophic and photoautotrophic) and aerobic chemoheterotrophy in the absence of light. [1] R. sphaeroides is also able to fix nitrogen. [2] It is remarkably metabolically diverse, as it is able to grow heterotrophically via fermentation and aerobic and anaerobic respiration. Such a metabolic versatility has motivated the investigation of R. sphaeroides as a microbial cell factory for biotechnological applications. [3]
Rhodobacter sphaeroides has been isolated from deep lakes and stagnant waters. [2]
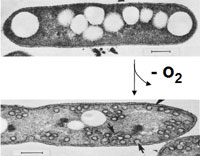
Rhodobacter sphaeroides is one of the most pivotal organisms in the study of bacterial photosynthesis. It requires no unusual conditions for growth and is incredibly efficient. The regulation of its photosynthetic machinery is of great interest to researchers, as R. sphaeroides has an intricate system for sensing O2 tensions. [4] Also, when exposed to a reduction in the partial pressure of oxygen, R. sphaeroides develops invaginations in its cellular membrane. The photosynthetic apparatus is housed in these invaginations. [4] These invaginations are also known as chromatophores.
The genome of R. sphaeroides is also somewhat intriguing. It has two chromosomes, one of 3 Mb (CI) and one of 900 Kb (CII), and five naturally occurring plasmids. Many genes are duplicated between the two chromosomes but appear to be differentially regulated. Moreover, many of the open reading frames (ORFs) on CII seem to code for proteins of unknown function. When genes of unknown function on CII are disrupted, many types of auxotrophy result, emphasizing that the CII is not merely a truncated version of CI. [5]
Bacterial small RNAs have been identified as components of many regulatory networks. Twenty sRNAs were experimentally identified in Rhodobacter spheroides, and the abundant ones were shown to be affected by singlet oxygen (1O2) exposure. [6] 1O2 which generates photooxidative stress, is made by bacteriochlorophyll upon exposure to oxygen and light. One of the 1O2 induced sRNAs SorY (1O2 resistance RNA Y) was shown to be induced under several stress conditions and conferred resistance against 1O2 by affecting a metabolite transporter. [7] SorX is the second 1O2 induced sRNA that counteracts oxidative stress by targeting mRNA for a transporter. It also has an impact on resistance against organic hydroperoxides. [8] A cluster of four homologous sRNAs called CcsR for conserved CCUCCUCCC motif stress-induced RNA has been shown to play a role in photo-oxidative stress resistance as well. [9] PcrZ (photosynthesis control RNA Z) identified in R. sphaeroides, is a trans-acting sRNA which counteracts the redox-dependent induction of photosynthesis genes, mediated by protein regulators. [10]
R. sphaeroides encodes several terminal oxidases which allow electron transfer to oxygen and other electron acceptors (e.g. DMSO or TMAO). [11] Therefore, this microorganism can respire under oxic, micro-oxic and anoxic conditions under both light and dark conditions. Moreover, it is capable to accept a variety of carbon substrates, including C1 to C4 molecules, sugars and fatty acids. [12] Several pathways for glucose catabolism are present in its genome, such as the Embden–Meyerhof–Parnas pathway (EMP), the Entner–Doudoroff pathway (ED) and the Pentose phosphate pathway (PP). [13] The ED pathway is the predominant glycolytic pathway in this microorganism, [14] whereas the EMP pathway contributing only to a smaller extent. [15] Variation in nutrient availability has important effects on the physiology of this bacterium. For example, decrease in oxygen tensions activates the synthesis of photosynthetic machinery (including photosystems, antenna complexes and pigments). Moreover, depletion of nitrogen in the medium triggers intracellular accumulation of polyhydroxybutyrate, a reserve polymer. [16]
A genome-scale metabolic model exists for this microorganism, [17] which can be used for predicting the effect of gene manipulations on its metabolic fluxes. For facilitating genome editing in this species, a CRISPR/Cas9 genome editing tool was developed [18] and expanded. [19] Moreover, partitioning of intracellular fluxes has been studied in detail, also with the help of 13C-glucose isotopomers. [15] [20] Altogether, these tools can be employed for improving R. sphaeroides as cell factory for industrial biotechnology. [3]
Knowledge of the physiology of R. sphaeroides allowed the development of biotechnological processes for the production of some endogenous compounds. These are hydrogen, polyhydroxybutyrate and isoprenoids (e.g. coenzyme Q10 and carotenoids). Moreover, this microorganism is used also for wastewater treatment. Hydrogen evolution occurs via the activity of the enzyme nitrogenase, [21] whereas isoprenoids are synthesized naturally via the endogenous MEP pathway. The native pathway has been optimized via genetic engineering for improving coenzyme Q10 synthesis. [22] Alternatively, improvement of isoprenoid synthesis was obtained via the introduction of a heterologous mevalonate pathway. [23] [16] Synthetic biology-driven engineering of the metabolism of R. sphaeroides, in combination to the functional replacement the MEP pathway with mevalonate pathway, [24] allowed to further increase bioproduction of isoprenoids in this species. [25]
- Rhodobacter sphaeroides (van Niel 1944) Imhoff et al., 1984 [26]
- Rhodococcus minor Molisch 1907
- Rhodococcus capsulatus Molisch 1907
- Rhodosphaera capsulata (Molisch) Buchanan 1918
- Rhodosphaera minor (Molisch) Bergey et al. 1923
- Rhodorrhagus minor (Molisch) Bergey et al. 1925
- Rhodorrhagus capsulatus (Molisch) Bergey et al. 1925
- Rhodorrhagus capsulatus Bergey et al. 1939
- Rhodopseudomonas sphaeroides van Niel 1944
- Rhodopseudomonas spheroides van Niel 1944
- Rhodorrhagus spheroides (van Niel) Brisou 1955
In 2020 it was recommended that Rhodobacter sphaeroides be moved to the genus Cereibacter. [27] This is the name currently used by the NCBI taxonomy database.
- ^ Mackenzie C, Eraso JM, Choudhary M, Roh JH, Zeng X, Bruscella P, et al. (2007). "Postgenomic adventures with Rhodobacter sphaeroides". Annu Rev Microbiol. 61: 283–307. doi: 10.1146/annurev.micro.61.080706.093402. PMID 17506668.
- ^ a b De Universiteit van Texas over Rhodobacter sphaeroides Archived 2009-07-10 at the Wayback Machine
- ^ a b Orsi E, Beekwilder J, Eggink G, Kengen SW, Weusthuis RA (2020). "The transition of Rhodobacter sphaeroides into a microbial cell factory". Biotechnology and Bioengineering. 118 (2): 531–541. doi: 10.1002/bit.27593. PMC 7894463. PMID 33038009.
- ^ a b Oh, JI.; Kaplan, S. (Mar 2001). "Generalized approach to the regulation and integration of gene expression". Mol Microbiol. 39 (5): 1116–23. doi: 10.1111/j.1365-2958.2001.02299.x. PMID 11251830. S2CID 27053575.
- ^ Mackenzie, C; Simmons, AE; Kaplan, S (1999). "Multiple chromosomes in bacteria. The yin and yang of trp gene localization in Rhodobacter sphaeroides 2.4.1". Genetics. 153 (2): 525–38. doi: 10.1093/genetics/153.2.525. PMC 1460784. PMID 10511537.
- ^ Berghoff, Bork A.; Glaeser, Jens; Sharma, Cynthia M.; Vogel, Jörg; Klug, Gabriele (2009-12-01). "Photooxidative stress-induced and abundant small RNAs in Rhodobacter sphaeroides". Molecular Microbiology. 74 (6): 1497–1512. doi: 10.1111/j.1365-2958.2009.06949.x. ISSN 1365-2958. PMID 19906181.
- ^ Adnan, Fazal; Weber, Lennart; Klug, Gabriele (2015-01-01). "The sRNA SorY confers resistance during photooxidative stress by affecting a metabolite transporter in Rhodobacter sphaeroides". RNA Biology. 12 (5): 569–577. doi: 10.1080/15476286.2015.1031948. ISSN 1555-8584. PMC 4615379. PMID 25833751.
- ^ Peng, Tao; Berghoff, Bork A.; Oh, Jeong-Il; Weber, Lennart; Schirmer, Jasmin; Schwarz, Johannes; Glaeser, Jens; Klug, Gabriele (2016-10-02). "Regulation of a polyamine transporter by the conserved 3' UTR-derived sRNA SorX confers resistance to singlet oxygen and organic hydroperoxides in Rhodobacter sphaeroides". RNA Biology. 13 (10): 988–999. doi: 10.1080/15476286.2016.1212152. ISSN 1555-8584. PMC 5056773. PMID 27420112.
- ^ Billenkamp, Fabian; Peng, Tao; Berghoff, Bork A.; Klug, Gabriele (May 2015). "A cluster of four homologous small RNAs modulates C1 metabolism and the pyruvate dehydrogenase complex in Rhodobacter sphaeroides under various stress conditions". Journal of Bacteriology. 197 (10): 1839–1852. doi: 10.1128/JB.02475-14. ISSN 1098-5530. PMC 4402390. PMID 25777678.
- ^ Mank, Nils N.; Berghoff, Bork A.; Hermanns, Yannick N.; Klug, Gabriele (2012-10-02). "Regulation of bacterial photosynthesis genes by the small noncoding RNA PcrZ". Proceedings of the National Academy of Sciences of the United States of America. 109 (40): 16306–16311. Bibcode: 2012PNAS..10916306M. doi: 10.1073/pnas.1207067109. ISSN 1091-6490. PMC 3479615. PMID 22988125.
- ^ Zannoni D, Schoepp-Cothenet B, Hosler J (2013). "Respiration and Respiratory Complexes". The Purple Phototrophic Bacteria. 28: 537–561. doi: 10.1186/1752-0509-7-89. ISSN 1752-0509. PMC 3849096. PMID 24034347.
- ^ Tabita FR (2004). "The Biochemistry and Metabolic Regulation of Carbon Metabolism and CO2 Fixation in Purple Bacteria". Anoxygenic Photosynthetic Bacteria. Advances in Photosynthesis and Respiration. 2: 885–914. doi: 10.1007/0-306-47954-0_41. ISBN 0-7923-3681-X.
- ^ Imam S, Noguera D, Donohue T. "Global insights into energetic and metabolic networks in Rhodobacter sphaeroides". BMC Systems Biology. 7 (1): 89. doi: 10.1007/0-306-47954-0_41.
- ^ Fuhrer T, Fischer E, Sauer U (2005). "Experimental Identification and Quantification of Glucose Metabolism in Seven Bacterial Species". Journal of Bacteriology. 187 (5): 1581–1590. doi: 10.1128/JB.187.5.1581-1590.2005. ISSN 0021-9193. PMC 1064017. PMID 15716428.
- ^ a b Orsi E, Beekwilder J, Peek S, Eggink G, Kengen SW, Weusthuis RA (2020). "Metabolic flux ratio analysis by parallel 13C labeling of isoprenoid biosynthesis in Rhodobacter sphaeroides". Metabolic Engineering. 57: 228–238. doi: 10.1016/j.ymben.2019.12.004. PMID 31843486.
- ^ a b Orsi E, Folch PL, Monje-Lopez V, Fernhout BM, Turcato A, Kengen SW, Eggink G, Weusthuis RA (2019). "Characterization of heterotrophic growth and sesquiterpene production by Rhodobacter sphaeroides on a defined medium". Journal of Industrial Microbiology & Biotechnology. 46 (8): 1179–1190. doi: 10.1007/s10295-019-02201-6. PMC 6697705. PMID 31187318.
- ^ Imam S, Ylmaz S, Sohmen U, Gorzalski AS, Reed JL, Noguera DR, Donohue TJ (2011). "iRsp1095: A genome-scale reconstruction of the Rhodobacter sphaeroides metabolic network". BMC Systems Biology. 5: 116. doi: 10.1186/1752-0509-5-116. PMC 3152904. PMID 21777427.
- ^ Mougiakos I, Orsi E, Rifqi-Ghiffary M, Post W, De Maria A, Adiego-Perez B, Kengen SW, Weusthuis RA, van der Oost J (2019). "Efficient Cas9-based genome editing of Rhodobacter sphaeroides for metabolic engineering". Microbial Cell Factories. 18 (1): 204. doi: 10.1186/s12934-019-1255-1. PMC 6876111. PMID 31767004.
- ^ Luo Y, Ge M, Wang B, Sun C, Wang J, Dong Y, Xi JJ (2020). "CRISPR/Cas9-deaminase enables robust base editing in Rhodobacter sphaeroides 2.4.1". Microbial Cell Factories. 19 (1): 93. doi: 10.1186/s12934-020-01345-w. PMC 7183636. PMID 32334589.
- ^ Tao Y, Liu D, Yan X, Zhou Z, Lee JK, Yang C (2012). "Network Identification and Flux Quantification of Glucose Metabolism in Rhodobacter sphaeroides under Photoheterotrophic H2-Producing Conditions". Journal of Bacteriology. 194 (2): 274–283. doi: 10.1128/JB.05624-11. PMC 3256653. PMID 22056932.
- ^ Luo Y, Ge M, Wang B, Sun C, Wang J, Dong Y, Xi JJ (2002). "Aspects of the metabolism of hydrogen production by Rhodobacter sphaeroides". International Journal of Hydrogen Energy. 27 (11–12): 1315–1329. doi: 10.1016/S0360-3199(02)00127-1.
- ^ Lu W, Ye L, Xu H, Xe W, Gu J, Yu H (2013). "Enhanced production of coenzyme Q10 by self‐regulating the engineered MEP pathway in Rhodobacter sphaeroides". Biotechnology and Bioengineering. 111 (4): 761–769. doi: 10.1002/bit.25130. PMID 24122603. S2CID 205503575.
- ^ Beekwilder J, van Houwelingen A, Cankar K, van Dijk AD, de Jong RM, Stoopen G, Bouwmeester H, Achkar J, Sonke T, Bosch D (2013). "Valencene synthase from the heartwook of Nootka cyperss (Callitropsis nootkatensis) for biotechnological production of valencene". BMC Systems Biology. 12 (2): 174–182. doi: 10.1111/pbi.12124. PMID 24112147.
- ^ Orsi E, Beekwilder J, van Gelder D, van Houwelingen A, Eggink G, Kengen SW, Weusthuis RA (2020). "Functional replacement of isoprenoid pathways in Rhodobacter sphaeroides". Microbial Biotechnology. 13 (4): 1082–1093. doi: 10.1111/1751-7915.13562. PMC 7264872. PMID 32207882.
- ^ Orsi E, Mougiakos I, Post W, Beekwilder J, Dompe M, Eggink G, van der Oost J, Kengen SW, Weusthuis RA (2020). "Growth-uncoupled isoprenoid synthesis in Rhodobacter sphaeroides". Biotechnology for Biofuels. 13: 123. doi: 10.1186/s13068-020-01765-1. PMC 7359475. PMID 32684976.
- ^ Bacteriology Insight Orienting System over Rhodobacter sphaeroides[ permanent dead link]
- ^ Hördt A, López MG, Meier-Kolthoff JP, Schleuning M, Weinhold LM, Tindall BJ, Gronow S, Kyrpides NC, Woyke T, Göker M (2020). "Analysis of 1,000+ Type-Strain Genomes Substantially Improves Taxonomic Classification of Alphaproteobacteria". Frontiers in Microbiology. 11: 468. doi: 10.3389/fmicb.2020.00468. PMC 7179689. PMID 32373076.
- Inomata Tsuyako, Higuchi Masataka (1976), Incorporation of tritium into cell materials of Rhodpseudomonas spheroides from tritiated water in the medium under aerobic conditions ; Journal of Biochemistry 80(3), p569-578, 1976-09