| RTL6 | |||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Identifiers | |||||||||||||||||||||||||||||||||||||||||||||||||||
| Aliases | RTL6, Mar6, Mart6, dJ1033E15.2, LDOC1L, leucine zipper, down-regulated in cancer 1-like, leucine zipper, down-regulated in cancer 1 like, LDOC1 like, SIRH3, retrotransposon Gag like 6 | ||||||||||||||||||||||||||||||||||||||||||||||||||
| External IDs | MGI: 2675858; HomoloGene: 18594; GeneCards: RTL6; OMA: RTL6 - orthologs | ||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| Wikidata | |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
Retrotransposon Gag Like 6 is a protein encoded by the RTL6 gene in humans. [5] RTL6 is a member of the Mart family of genes, which are related to Sushi-like retrotransposons and were derived from fish and amphibians. [6] The RTL6 protein is localized to the nucleus and has a predicted leucine zipper motif that is known to bind nucleic acids in similar proteins, such as LDOC1.
Gene
Locus
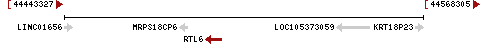
The gene is on Chromosome 22 (human) at 22q13.31 on the minus strand from 44492570 to 44498125 nt on the GRCh38.p7 assembly of the human genome. Aliases for the gene include LDOC1L, MAR6, MART6, and SIRH3. RTL6 is made up of 2 exons and is encoded by 5556 base pairs of DNA . [7]
Origin
RTL6 is a retrotransposon GAG related gene. It is one of eleven MART (Mammalian Retrotransposon Derived) genes in humans related to Sushi-like retrotransposons with long terminal repeats from fish and amphibians. [6] Between 170 and 310 MYA, MART genes lost their ability to retrotranspose and concomitantly gained new, beneficial function for its host organism. [8]
mRNA
RTL6 has an alternate start of transcription 140 base pairs upstream of the normal transcribed region. The lengths of the primary mRNA and that with the upstream start of transcription are 5355 and 5495 base pairs respectively. [7]
Protein
Primary information
The primary amino acid sequence for RTL6 is made up of 239 residues. [5] There are no known alternative splice variants of the protein. The molecular weight of the protein is 26.2 kDa and the isoelectric point is 11.58. [9] RTL6 is a proline and arginine rich protein. [9]


Domains and motifs
RTL6 contains a predicted leucine zipper motif known to participate in nucleic acid binding in other proteins. [9] RTL6 also contains a domain of unknown function from amino acid residues 98-177 . RTL6 is one of a number of genes belonging to the DUF4939 (domain of unknown function) superfamily. [14]
Secondary structure
The secondary structure of RTL6 is made up of largely alpha helices. [15] One region of RTL6 is also predicted to participate in a coiled-coil structure from amino acid residues 29–63. [14]
Post-translational modifications
There are also two predicted phosphorylation sites for Protein Kinase C with high confidence scores at amino acid residues 6 and 45. [16] [17] There is also a predicted ubiquitination site with medium-confidence at amino acid residue 8. [18]

Cellular sublocation
RTL6 is expected to be localized to the nucleus and cytosol based on the presence of a leucine zipper domain, the absence of signals indicating secretion or transmembrane domains, and immunohistochemical staining. [19] [20] [21]
Expression
RTL6 has been shown to be expressed at high levels during all stages of development and in a wide variety of tissues. [22] [23] [13]
RTL6 expression has been shown to fall in HeLa cervical cancer cells upon treatment with chemotherapeutic Casiopeinas and in A549 lung cancer cells upon treatment with Actinomycin D. [24] [25]
Interacting proteins
RTL6 has been shown to interact with the following proteins:
| DDIT3 | DNA damage-inducible transcript 3 protein [26] |
| NXF1 | Nuclear RNA export factor 1 [27] |
| STX18 | Syntaxin 18 [28] |
| MAFF | MAF bZIP transcription factor F [28] |
| GOPC | Golgi-associated PDZ and coiled-coil motif-containing protein [28] |
| BATF3 | Basic leucine zipper transcriptional factor ATF-like 3 [28] |
| TERF2 | Telomeric repeat-binding factor 2 [29] |
| UXAC | Uronate isomerase ( Yersinia pestis) [30] |
Clinical significance
The RTL6 protein has been shown to interact with the UXAC protein from Yersinia pestis, the gram-negative bacterium responsible for the bubonic plague. [30]
Homology/evolution
Paralogs
Eleven paralogs were identified for RTL6 in humans. The paralogs have diverse functions and expression patterns, although many are known to have zinc finger domains and bind nucleic acids:
| MART Family Name | Accession Number | Sequence Length | Query Cover | Percent Similarity |
|---|---|---|---|---|
| RTL1 [31] | NP_001128360.1 | 1358 | 100 | 7.6 |
| PEG10 (RTL2) [32] | NP_055883.2 | 359 | 35 | 35 |
| RTL3 [33] | NP_689907.1 | 2648 | 100 | 2.2 |
| RTL4 [34] | NP_001004308.2 | 310 | 100 | 19 |
| RTL5 [35] | NP_001019626.1 | 569 | 37 | 53 |
| RTL6 [5] | NP_115663.2 | 239 | NA | NA |
| LDOC1 (RTL7) [36] | NP_036449.1 | 146 | 33 | 41 |
| RTL8A [37] | NP_001071640.1 | 113 | 33 | 44 |
| RTL8B [38] | NP_001071641.1 | 113 | 33 | 43 |
| RTL8C [39] | NP_001071639.1 | 113 | 33 | 44 |
| RTL9 [40] | NP_065820.1 | 1388 | 22 | 39 |
| RTL10 [41] | NP_078903.3 | 364 | 34 | 35 |
Orthologs
RTL6 is highly conserved across mammals, including the leucine zipper motif and DUF4939. The gene is also conserved in marsupials such as the opossum but not in birds such as the chicken, suggesting the gene was likely formed after the divergence of mammals and birds but before the divergence of marsupials and mammals (170-310 MYA: [6]
| Organism | Common Name | Classification | Accession Number | Percent Identity | Query Cover | Percent Similarity |
|---|---|---|---|---|---|---|
| Homo sapiens | Humans | Primate | NP_115663.2 [5] | NA | NA | NA |
| Macaca mulatta | Rhesus Monkey | Primate | NP_001181372.1 [42] | 98 | 100 | 99 |
| Felis catus | House Cat | Carnivore | XP_003989415.1 [43] | 97 | 100 | 98 |
| Mus muscalus | Common Mouse | Rodent | NP_808298.2 [44] | 92 | 100 | 96 |
| Pteropus alecto | Black Flying Fox | Bat | XP_006917396.1 [45] | 96 | 100 | 98 |
| Equus caballus | Horse | Odd-Toed Ungulates | XP_005606827.1 [46] | 95 | 100 | 98 |
| Bos Taurus | Cattle | Even-Toed Ungulates | XP_015326927.1 [47] | 94 | 100 | 97 |
| Orcinus orca | Killer Whale | Whales/Dolphins | XP_004279624.1 [48] | 95 | 100 | 98 |
| Trichechus manatus latirostirs | Florida Manatee | Placentals | XP_004380056.1 [49] | 95 | 100 | 96 |
| Erinaceus europaeus | European Hedgehog | Rabbits/Hares | XP_016043235.1 [50] | 93 | 100 | 96 |
| Ochotona princeps | American Pika | Insectivoires | XP_004589491.1 [51] | 92 | 100 | 97 |
The most distantly detectable organisms with homology in the gene are bony fishes including salmon and the common carp, but similarity to the human protein sequence is markedly less than that of mammals. No traces of the gene can be seen in intermediates between mammals and bony fishes such as reptiles or amphibians:
| Organism | Common Name | Classification | Accession Number | Percent Identity | Query Cover | Percent Similarity |
|---|---|---|---|---|---|---|
| Homo sapiens | Humans | Primate | NP_115663.2 [5] | 100 | 100 | 100 |
| Cyprinus carpio | Common Carp | Bony Fishes | XP_018946777 [52] | 30 | 41 | 34 |
| Esox lucius | Northern Pike | Bony Fishes | XP_019899574.1 [53] | 31 | 55 | 31 |
| Nothobranchius furzeri | Black Rockcod | Bony Fishes | XP_010767110 [54] | 37 | 35 | 39 |
References
- ^ a b c GRCh38: Ensembl release 89: ENSG00000188636 – Ensembl, May 2017
- ^ a b c GRCm38: Ensembl release 89: ENSMUSG00000055745 – Ensembl, May 2017
- ^ "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ a b c d e f "RTL6 retrotransposon Gag like 6 [Homo sapiens (human)] - Gene - NCBI". www.ncbi.nlm.nih.gov. Retrieved 2017-05-02.
- ^ a b c Brandt J, Schrauth S, Veith AM, Froschauer A, Haneke T, Schultheis C, Gessler M, Leimeister C, Volff JN (January 2005). "Transposable elements as a source of genetic innovation: expression and evolution of a family of retrotransposon-derived neogenes in mammals". Gene. 345 (1): 101–11. doi: 10.1016/j.gene.2004.11.022. PMID 15716091.
- ^ a b "LDOC1L LDOC1 like [Homo sapiens (human)] - Gene - NCBI". www.ncbi.nlm.nih.gov. Retrieved 2017-02-05.
- ^ Brandt J, Veith AM, Volff JN (2005-01-01). "A family of neofunctionalized Ty3/gypsy retrotransposon genes in mammalian genomes". Cytogenetic and Genome Research. 110 (1–4): 307–17. doi: 10.1159/000084963. PMID 16093683. S2CID 38398479.
- ^ a b c Brendel, V., Bucher, P., Nourbakhsh, I.R., Blaisdell, B.E. & Karlin, S. (1992) "Methods and algorithms for statistical analysis of protein sequences" Proc. Natl. Acad. Sci. U.S.A. 89, 2002-2006.
- ^ J Yang, R Yan, A Roy, D Xu, J Poisson, Y Zhang. The I-TASSER Suite: Protein structure and function prediction. Nature Methods, 12: 7-8, 2015.
- ^ A Roy, A Kucukural, Y Zhang. I-TASSER: a unified platform for automated protein structure and function prediction. Nature Protocols, 5: 725-738, 2010.
- ^ Y Zhang. I-TASSER server for protein 3D structure prediction. BMC Bioinformatics, 9: 40, 2008.
- ^ a b "Cell atlas - LDOC1L - The Human Protein Atlas". www.proteinatlas.org. Retrieved 2017-05-02.
- ^ a b "LDOC1L - Protein LDOC1L - Homo sapiens (Human) - LDOC1L gene & protein". www.uniprot.org. Retrieved 2017-05-07.
- ^ Garnier, Gibrat, and Robson, Meth. Enzymol., R.F. Doolittle ed. (1996) 266:97-120
- ^ "Motif Scan". myhits.isb-sib.ch. Retrieved 2017-05-02.
- ^ Blom N, Gammeltoft S, Brunak S (December 1999). "Sequence and structure-based prediction of eukaryotic protein phosphorylation sites". Journal of Molecular Biology. 294 (5): 1351–62. doi: 10.1006/jmbi.1999.3310. PMID 10600390.
- ^ Radivojac P, Vacic V, Haynes C, Cocklin RR, Mohan A, Heyen JW, Goebl MG, Iakoucheva LM (February 2010). "Identification, analysis, and prediction of protein ubiquitination sites". Proteins. 78 (2): 365–80. doi: 10.1002/prot.22555. PMC 3006176. PMID 19722269.
- ^ "PSORT: Protein Subcellular Localization Prediction Tool". www.genscript.com. Retrieved 2017-05-02.
- ^ Lin WZ, Fang JA, Xiao X, Chou KC (April 2013). "iLoc-Animal: a multi-label learning classifier for predicting subcellular localization of animal proteins". Molecular BioSystems. 9 (4): 634–44. doi: 10.1039/c3mb25466f. PMID 23370050.
- ^ "BLAST: Basic Local Alignment Search Tool". blast.ncbi.nlm.nih.gov. Retrieved 2017-02-19.
- ^ "Gene Detail :: Allen Brain Atlas: Mouse Brain". mouse.brain-map.org. Retrieved 2017-05-02.
- ^ "Leucine zipper, down-regulated in cancer 1-like (Ldoc1l)". www.ncbi.nlm.nih.gov. Retrieved 2017-05-02.
- ^ Serment-Guerrero J, Cano-Sanchez P, Reyes-Perez E, Velazquez-Garcia F, Bravo-Gomez ME, Ruiz-Azuara L (October 2011). "Genotoxicity of the copper antineoplastic coordination complexes casiopeinas". Toxicology in Vitro. 25 (7): 1376–84. doi: 10.1016/j.tiv.2011.05.008. PMID 21601632.
- ^ Capranico G, Binaschi M (October 1998). "DNA sequence selectivity of topoisomerases and topoisomerase poisons". Biochimica et Biophysica Acta (BBA) - Gene Structure and Expression. 1400 (1–3): 185–94. doi: 10.1016/S0167-4781(98)00135-3. PMID 9748568.
- ^ Behrends C, Sowa ME, Gygi SP, Harper JW (July 2010). "Network organization of the human autophagy system". Nature. 466 (7302): 68–76. Bibcode: 2010Natur.466...68B. doi: 10.1038/nature09204. PMC 2901998. PMID 20562859.
- ^ Castello A, Fischer B, Eichelbaum K, Horos R, Beckmann BM, Strein C, Davey NE, Humphreys DT, Preiss T, Steinmetz LM, Krijgsveld J, Hentze MW (June 2012). "Insights into RNA biology from an atlas of mammalian mRNA-binding proteins". Cell. 149 (6): 1393–406. doi: 10.1016/j.cell.2012.04.031. PMID 22658674.
- ^ a b c d Huttlin EL, Ting L, Bruckner RJ, Gebreab F, Gygi MP, Szpyt J, Tam S, Zarraga G, Colby G, Baltier K, Dong R, Guarani V, Vaites LP, Ordureau A, Rad R, Erickson BK, Wühr M, Chick J, Zhai B, Kolippakkam D, Mintseris J, Obar RA, Harris T, Artavanis-Tsakonas S, Sowa ME, De Camilli P, Paulo JA, Harper JW, Gygi SP (July 2015). "The BioPlex Network: A Systematic Exploration of the Human Interactome". Cell. 162 (2): 425–440. doi: 10.1016/j.cell.2015.06.043. PMC 4617211. PMID 26186194.
- ^ Giannone RJ, McDonald HW, Hurst GB, Shen RF, Wang Y, Liu Y (August 2010). "The protein network surrounding the human telomere repeat binding factors TRF1, TRF2, and POT1". PLOS ONE. 5 (8): e12407. Bibcode: 2010PLoSO...512407G. doi: 10.1371/journal.pone.0012407. PMC 2928292. PMID 20811636.
- ^ a b Dyer MD, Neff C, Dufford M, Rivera CG, Shattuck D, Bassaganya-Riera J, Murali TM, Sobral BW (August 2010). "The human-bacterial pathogen protein interaction networks of Bacillus anthracis, Francisella tularensis, and Yersinia pestis". PLOS ONE. 5 (8): e12089. Bibcode: 2010PLoSO...512089D. doi: 10.1371/journal.pone.0012089. PMC 2918508. PMID 20711500.
- ^ "RTL1 retrotransposon Gag like 1 [Homo sapiens (human)] - Gene - NCBI". www.ncbi.nlm.nih.gov. Retrieved 2017-05-06.
- ^ "PEG10 paternally expressed 10 [Homo sapiens (human)] - Gene - NCBI". www.ncbi.nlm.nih.gov. Retrieved 2017-05-06.
- ^ "RTL3 retrotransposon Gag like 3 [Homo sapiens (human)] - Gene - NCBI". www.ncbi.nlm.nih.gov. Retrieved 2017-05-06.
- ^ "RTL4 retrotransposon Gag like 4 [Homo sapiens (human)] - Gene - NCBI". www.ncbi.nlm.nih.gov. Retrieved 2017-05-06.
- ^ "RTL5 retrotransposon Gag like 5 [Homo sapiens (human)] - Gene - NCBI". www.ncbi.nlm.nih.gov. Retrieved 2017-05-06.
- ^ "LDOC1 LDOC1, regulator of NFKB signaling [Homo sapiens (human)] - Gene - NCBI". www.ncbi.nlm.nih.gov. Retrieved 2017-05-06.
- ^ "RTL8A retrotransposon Gag like 8A [Homo sapiens (human)] - Gene - NCBI". www.ncbi.nlm.nih.gov. Retrieved 2017-05-06.
- ^ "RTL8B retrotransposon Gag like 8B [Homo sapiens (human)] - Gene - NCBI". www.ncbi.nlm.nih.gov. Retrieved 2017-05-06.
- ^ "RTL8C retrotransposon Gag like 8C [Homo sapiens (human)] - Gene - NCBI". www.ncbi.nlm.nih.gov. Retrieved 2017-05-06.
- ^ "RTL9 retrotransposon Gag like 9 [Homo sapiens (human)] - Gene - NCBI". www.ncbi.nlm.nih.gov. Retrieved 2017-05-06.
- ^ "RTL10 retrotransposon Gag like 10 [Homo sapiens (human)] - Gene - NCBI". www.ncbi.nlm.nih.gov. Retrieved 2017-05-06.
- ^ "RTL6 retrotransposon Gag like 6 [Macaca mulatta (Rhesus monkey)] - Gene - NCBI". www.ncbi.nlm.nih.gov. Retrieved 2017-05-06.
- ^ "RTL6 retrotransposon Gag like 6 [Felis catus (domestic cat)] - Gene - NCBI". www.ncbi.nlm.nih.gov. Retrieved 2017-05-06.
- ^ "Ldoc1l leucine zipper, down-regulated in cancer 1-like [Mus musculus (house mouse)] - Gene - NCBI". www.ncbi.nlm.nih.gov. Retrieved 2017-05-06.
- ^ "RTL6 retrotransposon Gag like 6 [Pteropus alecto (black flying fox)] - Gene - NCBI". www.ncbi.nlm.nih.gov. Retrieved 2017-05-06.
- ^ "RTL6 retrotransposon Gag like 6 [Equus caballus (horse)] - Gene - NCBI". www.ncbi.nlm.nih.gov. Retrieved 2017-05-06.
- ^ "RTL6 retrotransposon Gag like 6 [Bos taurus (cattle)] - Gene - NCBI". www.ncbi.nlm.nih.gov. Retrieved 2017-05-06.
- ^ "RTL6 retrotransposon Gag like 6 [Orcinus orca (killer whale)] - Gene - NCBI". www.ncbi.nlm.nih.gov. Retrieved 2017-05-06.
- ^ "LOC101345980 retrotransposon Gag like 6 [Trichechus manatus latirostris (Florida manatee)] - Gene - NCBI". www.ncbi.nlm.nih.gov. Retrieved 2017-05-06.
- ^ "RTL6 retrotransposon Gag like 6 [Erinaceus europaeus (western European hedgehog)] - Gene - NCBI". www.ncbi.nlm.nih.gov. Retrieved 2017-05-06.
- ^ "RTL6 retrotransposon Gag like 6 [Ochotona princeps (American pika)] - Gene - NCBI". www.ncbi.nlm.nih.gov. Retrieved 2017-05-06.
- ^ "PREDICTED: protein LDOC1-like [Cyprinus carpio] - Protein - NCBI". www.ncbi.nlm.nih.gov. Retrieved 2017-05-06.
- ^ "PREDICTED: protein LDOC1L-like [Esox lucius] - Protein - NCBI". www.ncbi.nlm.nih.gov. Retrieved 2017-05-06.
- ^ "PREDICTED: uncharacterized protein LOC104943421 [Notothenia coriiceps] - Protein - NCBI". www.ncbi.nlm.nih.gov. Retrieved 2017-05-06.



