A peptide sequence tag is a piece of information about a peptide obtained by tandem mass spectrometry that can be used to identify this peptide in a protein database. [1] [2] [3]
Mass spectrometry
In general, peptides can be identified by fragmenting them in a mass spectrometer. For example, during collision-induced dissociation peptides collide with a gas within the mass spectrometer and break into pieces at their peptide bonds. The resulting fragment ions (called b-ions and y-ions) have mass differences corresponding to the residue masses of the respective amino acids. Thus, a tandem mass spectrum contains partial information about the amino acid sequence of the peptide. The peptide sequence tag approach, developed by Matthias Wilm and Matthias Mann at the EMBL, [4] uses this information to identify the peptide in a database. Briefly, a couple of masses are extracted from the spectrum in order to obtain the peptide sequence tag. This peptide sequence tag is a unique identifier of a specific peptide and can be used to find it in a database containing all possible peptide sequences.
Peptide fragment notation
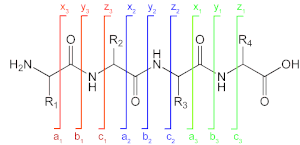
A notation has been developed for indicating peptide fragments that arise from a tandem mass spectrum. [5] Peptide fragment ions are indicated by a, b, or c if the charge is retained on the N-terminus and by x, y or z if the charge is maintained on the C-terminus. The subscript indicates the number of amino acid residues in the fragment. Prime symbols indicate the number of protons or hydrogens added to the fragment to form the observed ion. For example, y'' denotes the singly charged ion analogous to a protonated peptide, (y''')2+ is a doubly charged ion analogous to a doubly protonated peptide. [6]
See also
References
- ^ Hardouin J (2007). "Protein sequence information by matrix-assisted laser desorption/ionization in-source decay mass spectrometry". Mass Spectrometry Reviews. 26 (5): 672–82. Bibcode: 2007MSRv...26..672H. doi: 10.1002/mas.20142. PMID 17492750.
- ^ Shadforth I, Crowther D, Bessant C (2005). "Protein and peptide identification algorithms using MS for use in high-throughput, automated pipelines". Proteomics. 5 (16): 4082–95. doi: 10.1002/pmic.200402091. PMID 16196103. S2CID 38068737.
- ^ Mørtz E, O'Connor PB, Roepstorff P, Kelleher NL, Wood TD, McLafferty FW, Mann M (1996). "Sequence tag identification of intact proteins by matching tanden mass spectral data against sequence data bases". Proc. Natl. Acad. Sci. U.S.A. 93 (16): 8264–7. Bibcode: 1996PNAS...93.8264M. doi: 10.1073/pnas.93.16.8264. PMC 38658. PMID 8710858.
- ^ Mann M, Wilm M (1994). "Error-tolerant identification of peptides in sequence databases by peptide sequence tags". Anal. Chem. 66 (24): 4390–9. doi: 10.1021/ac00096a002. PMID 7847635.
- ^ a b Roepstorff P, Fohlman J (1984). "Proposal for a common nomenclature for sequence ions in mass spectra of peptides". Biomed. Mass Spectrom. 11 (11): 601. doi: 10.1002/bms.1200111109. PMID 6525415.
- ^ Tang XJ, Thibault P, Boyd RK (October 1993). "Fragmentation reactions of multiply protonated peptides and implications for sequencing by tandem mass spectrometry with low-energy collision-induced dissociation". Anal. Chem. 65 (20): 2824–34. doi: 10.1021/ac00068a020. PMID 7504416.