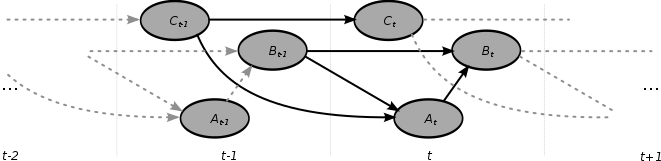
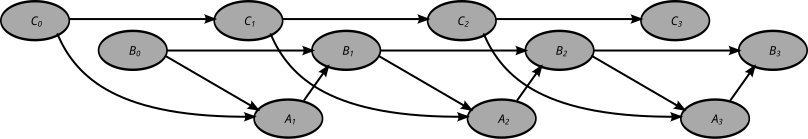

A dynamic Bayesian network (DBN) is a Bayesian network (BN) which relates variables to each other over adjacent time steps.
History
A dynamic Bayesian network (DBN) is often called a "two-timeslice" BN (2TBN) because it says that at any point in time T, the value of a variable can be calculated from the internal regressors and the immediate prior value (time T-1). DBNs were developed by Paul Dagum in the early 1990s at Stanford University's Section on Medical Informatics. [1] [2] Dagum developed DBNs to unify and extend traditional linear state-space models such as Kalman filters, linear and normal forecasting models such as ARMA and simple dependency models such as hidden Markov models into a general probabilistic representation and inference mechanism for arbitrary nonlinear and non-normal time-dependent domains. [3] [4]
Today, DBNs are common in robotics, and have shown potential for a wide range of data mining applications. For example, they have been used in speech recognition, digital forensics, protein sequencing, and bioinformatics. DBN is a generalization of hidden Markov models and Kalman filters. [5]
DBNs are conceptually related to probabilistic Boolean networks [6] and can, similarly, be used to model dynamical systems at steady-state.
See also
References
- ^ Paul Dagum; Adam Galper; Eric Horvitz (1992). "Dynamic Network Models for Forecasting" (PDF). Proceedings of the Eighth Conference on Uncertainty in Artificial Intelligence. AUAI Press: 41–48.
- ^ Paul Dagum; Adam Galper; Eric Horvitz; Adam Seiver (1995). "Uncertain Reasoning and Forecasting". International Journal of Forecasting. 11 (1): 73–87. doi: 10.1016/0169-2070(94)02009-e.
- ^ Paul Dagum; Adam Galper; Eric Horvitz (June 1991). "Temporal Probabilistic Reasoning: Dynamic Network Models for Forecasting" (PDF). Knowledge Systems Laboratory. Section on Medical Informatics, Stanford University.
- ^ Paul Dagum; Adam Galper; Eric Horvitz (1993). "Forecasting Sleep Apnea with Dynamic Network Models". Proceedings of the Ninth Conference on Uncertainty in Artificial Intelligence. AUAI Press: 64–71.
-
^
Stuart Russell;
Peter Norvig (2010).
Artificial Intelligence: A Modern Approach (PDF) (Third ed.).
Prentice Hall. p. 566.
ISBN
978-0136042594. Archived from
the original (PDF) on 20 October 2014. Retrieved 22 October 2014.
dynamic Bayesian networks (which include hidden Markov models and Kalman filters as special cases)
- ^ Harri Lähdesmäki; Sampsa Hautaniemi; Ilya Shmulevich; Olli Yli-Harja (2006). "Relationships between probabilistic Boolean networks and dynamic Bayesian networks as models of gene regulatory networks". Signal Processing. 86 (4): 814–834. doi: 10.1016/j.sigpro.2005.06.008. PMC 1847796. PMID 17415411.
Further reading
- Murphy, Kevin (2002). Dynamic Bayesian Networks: Representation, Inference and Learning. UC Berkeley, Computer Science Division.
- Ghahramani, Zoubin (1998). "Learning dynamic Bayesian networks". Adaptive Processing of Sequences and Data Structures. Lecture Notes in Computer Science. Vol. 1387. pp. 168–197. CiteSeerX 10.1.1.56.7874. doi: 10.1007/BFb0053999. ISBN 978-3-540-64341-8.
- Friedman, N.; Murphy, K.; Russell, S. (1998). Learning the structure of dynamic probabilistic networks. UAI’98. Morgan Kaufmann. pp. 139–147. CiteSeerX 10.1.1.75.2969.
- Shiguihara, P.; De Andrade Lopes, A.; Mauricio, D. (2021).
"Dynamic Bayesian Network Modeling, Learning, and Inference: A Survey". IEEE Access.
doi:
10.1109/ACCESS.2021.3105520.
{{ cite journal}}: Cite journal requires|journal=( help)
Software
- bnt on GitHub: the Bayes Net Toolbox for Matlab, by Kevin Murphy, (released under a GPL license)
- Graphical Models Toolkit (GMTK): an open-source, publicly available toolkit for rapidly prototyping statistical models using dynamic graphical models (DGMs) and dynamic Bayesian networks (DBNs). GMTK can be used for applications and research in speech and language processing, bioinformatics, activity recognition, and any time-series application.
- DBmcmc : Inferring Dynamic Bayesian Networks with MCMC, for Matlab (free software)
- GlobalMIT Matlab toolbox at Google Code: Modeling gene regulatory network via global optimization of dynamic bayesian network (released under a GPL license)
- libDAI: C++ library that provides implementations of various (approximate) inference methods for discrete graphical models; supports arbitrary factor graphs with discrete variables, including discrete Markov Random Fields and Bayesian Networks (released under the FreeBSD license)
- aGrUM: C++ library (with Python bindings) for different types of PGMs including Bayesian Networks and Dynamic Bayesian Networks (released under the GPLv3)
- FALCON: Matlab toolbox for contextualization of DBNs models of regulatory networks with biological quantitative data, including various regularization schemes to model prior biological knowledge (released under the GPLv3)