An Artificial Metalloenzyme (ArM) is a designer metalloprotein, not found in nature, which can catalyze desired chemical reactions. [1] [2] Despite fitting into classical enzyme categories, ArMs also have potential in new-to-nature chemical reactivity like catalysing Suzuki coupling, [3] Metathesis [4] etc., which were never reported among natural enzymatic reactions.

ArMs have two main components: a protein scaffold and an artificial catalytic moiety, which, in this case, features a metal center. This class of designer biocatalysts is unique because of the potential to improve the catalytic performance through chemogenetic optimization, a parallel improvement of both the direct metal surrounding (first coordination sphere) and the protein scaffold (second coordination sphere).The second coordination sphere (protein scaffold) is easily evolvable and, in the case of ArMs, responsible for very high (stereo)selectivity. [5] With the progress in organometallic synthesis and protein engineering, more and more new kind of design of ArMs were developed, showing promising future in both academia and industrial aspects. [6]
In 2018, one-half of the Nobel Prize in Chemistry was awarded to Frances H. Arnold "for the directed evolution of enzymes", who elegantly evolved artificial metalloenzymes to realize efficient and highly selective new-to-nature chemical reactions in vitro and in vivo.
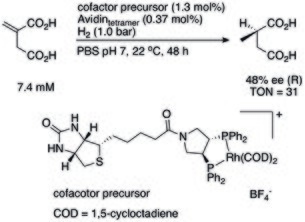
Dated back to 1956, the first protein modified transition metal catalyst was documented. [7] The Palladium(II) salt was absorbed onto silk fibroin fiber, reduced by hydrogen to get the first reported ArM, which can catalyze asymmetric hydrogenation. This work was not reproducible, but it is considered to be the first work in the field of artificial metalloenzymes. [5] At that time, the major challenge that blocked further studies was underdeveloped protein production and purification technology. The first attempt to anchor an abiotic metal center onto a protein was reported by Whitesides et al. using biotin- avidin interaction, making an artificial hydrogenase. [8] The presence of avidin can significantly increase the catalytic capacity of Rhodium(I) cofactor in aqueous phosphate buffer. Another pioneering work was conducted by Kaiser et al. where carboxypeptidase A (CPA) was repurposed into an oxidase by substituting Zn(II) center by Cu(II), for the oxidation of ascorbic acid. [9]
The real potential of ArMs was unleashed when recombinant protein production was developed, namely in 1997 Distefano and Davies reported a scaffold modification of a recombinant adipocyte lipid-binding protein (ALBP) with iodoacetamido-1,10-phenanthroline coordinating Cu(II) for the stereoselective hydrolysis of racemic esters. [10]
Four strategies have been used to assemble ArMs: [6]
- Covalent immobilization of a metal-containing catalytic moiety by an irreversible reaction with the protein;
- Supramolecular interactions between a protein and a high-affinity substrate could be used to anchor a metal cofactor;
- The metal substitution in a natural metalloenzyme can result in a novel catalytic activity to the protein. The metal could be part of a prosthetic group (e.g., heme) or bound to amino acids;
- Amino acids with Lewis-basic properties in a hydrophobic pocket could interact with coordinatively unsaturated metal center.
These four strategies led to a great progress in the field of artificial metalloenzymes since the beginning of the 21st century, unlocking exceptional selectivity for new-to-nature reactions.

With the development of bioconjugation technology, there are plenty of strategies to covalently bind an artificial metallocofactor onto a protein scaffold:
- cysteine residue based chemistry: Cys- meleimide, [11] Cys- α-haloketone, [12] Cys-benzylhalide chemistry and disulfide formation, [13]
- post-translational bioorthogonal modification based on Amber stop codon suppression (e.g., Click chemistry) [14]
- enzyme active site modification (e.g., covalent bond formation between lipase and lipase inhibitor). [15]
Streptavidin or avidin in combination with biotinylated artificial metal cofactors is the most commonly used supramolecular strategy to make ArMs. [16] In the early example from Ward et al. shown below, the ligand of Ru(I) complex was covalently linked to biotin and than the whole complex was anchored to streptavidin thanks to a specific and strong biotin-streptavidin interaction. [17] The formed ArM can catalyze the reduction of prochiral ketones. Taking advantages of protein evolvability, different mutants of streptavidin can achieve different stereoselectivity. Throughout the years, many streptavidin-based enzymes were developed, enabling catalysis of very complex transformations in water, under ambient conditions.
Besides biotin-streptavidin based ArMs, another important example of using supramolecular iassembly strategy is antigen- antibody recognition. First reported in 1989 by Lerner et al.., a monoclonal antibody-based ArM is raised to hydrolyze specific peptide. [18]
Another interesting scaffold used as a platform for supramolecularly assembled ArMs are multidrug resistance regulators (MDRs), particularly a PadR family of proteins without native catalytic activity, whose function in nature is the recognition of foreign agents and to activate subsequent cellular response. [19] Among them, Lactococcal multidrug resistance regulator (LmrR) was mainly used to create ArMs, using different strategies, including the supramolecular one. Namely, Roelfes et al. incorporated Cu(II) phenanthroline complex in the hydrophobic pocket of LmrR and performed Friedel-Crafts reaction enantioselectively; [20] [21] and Fe heme complex which catalyzed cyclopropanation enantioselectively. [22]

This strategy involves substitution of a native metal center in a metallocofactor, by another metal, that might or might not be already present in living systems. [23] In this way, electronic and steric properties of the catalytic active site are altered compared to the wild-type enzyme, and novel catalytic pathways are unlocked.

The dative anchoring strategy uses natural amino acid residue in the protein scaffold like His, Cys, Glu, Asp and Ser to coordinate to a metal center. Like the first example of Pd-fibroin, dative anchoring to natural amino acids is not commonly used nowadays and often resulted in a more ambiguous binding site for metal compared with previous three methods.
However, these challenges can be overcome by in vivo incorporating metal-chelating non-canonical amino acids (ncAAs) [24] in the protein scaffold. These genetically encoded ncAAs' side chains have chelating moieties, such as 2,2'-bipyridine (3-(2,2'-bipyridin-5-yl)-L-alanine) [25] and 8-hydroxyquinoline (2-amino-3-(8-hydroxyquinolin-3-yl)propanoic acid) [26] that can selectively coordinate different metals. Combining protein scaffolds featuring chelating ncAAs with different metals yields exceptionally selective artificial metalloenzymes with various application potentials. [5] ncAAs are usually incorporated through the means of Amber stop codon suppression, via the orthogonal translation system (OTS). [24]
In addition to anchoring artificial metal center in the protein scaffold, researchers like Frances Arnold and Yang Yang focused on changing the native environment of natural metallocofactors. Due to the large sequence space that can be evolved in natural metalloenzymes, they can be evolved to catalyse non-native transformations. This process is known as enzyme repurposing. Directed evolution is commonly used to tailor the catalytic capacity and repurpose the enzyme function. Mostly based on native porphyrin-metallocofactor, Arnold's lab has developed many ArMs catalysing regioselective and/or enantioselective transformations, such as Carbon- Boron bond formation, [27] carbene insertion, [28] and aminohydroxylation [29] by evolving the sequence context of the corresponding ArMs.
As the pioneers of metalloredox radical biocatalysis, Yang et al. repurposed cytochrome P450s to catalyze atom transfer radical cyclization (ATRC), [30] and Huang et al. repurposed non-heme Fe-dependent enzymes to catalyze an abiological radical-relay azidation [31] and radical fluorination. [32] [33]
So far, ArMs can catalyze planty of chemical reactions, such as: allylic alkylation, allylic amination, aldol reaction, alcohol oxidation, C-H activation, [34] click reaction, [35] catechol oxidation, CO2 reduction, cyclopropanation, [36] Diels-Alder reaction, [37] epoxidation, epoxide ring opening, Friedel-Crafts alkylation, [38] hydrogenation, hydroformylation, Heck reaction, Metathesis, [4] Michael addition, nitrite reduction, NO reduction, Suzuki reaction, [3] Si-H insertion, [39] polymerization ( atom transfer radical polymerization), [40] atom transfer radical cyclization (ATRC) [30] and radical fluorination. [32] [33]
- ^ Morra S, Pordea A (October 2018). "Biocatalyst-artificial metalloenzyme cascade based on alcohol dehydrogenase". Chemical Science. 9 (38). Royal Society of Chemistry: 7447–7454. doi: 10.1039/C8SC02371A. PMC 6180310. PMID 30319745.
- ^ Leurs M, Dorn B, Wilhelm S, Manisegaran M, Tiller JC (July 2018). "Multicore Artificial Metalloenzymes Derived from Acylated Proteins as Catalysts for the Enantioselective Dihydroxylation and Epoxidation of Styrene Derivatives". Chemistry: A European Journal. 24 (42): 10859–10867. doi: 10.1002/chem.201802185. PMID 29808506.
- ^ a b Chatterjee A, Mallin H, Klehr J, Vallapurackal J, Finke AD, Vera L, et al. (January 2016). "An enantioselective artificial Suzukiase based on the biotin-streptavidin technology". Chemical Science. 7 (1): 673–677. doi: 10.1039/C5SC03116H. PMC 5953008. PMID 29896353.
- ^ a b Jeschek M, Reuter R, Heinisch T, Trindler C, Klehr J, Panke S, et al. (September 2016). "Directed evolution of artificial metalloenzymes for in vivo metathesis" (PDF). Nature. 537 (7622): 661–665. Bibcode: 2016Natur.537..661J. doi: 10.1038/nature19114. PMID 27571282. S2CID 205250261.
- ^ a b c Davis HJ, Ward TR (July 2019). "Artificial Metalloenzymes: Challenges and Opportunities". ACS Central Science. 5 (7): 1120–1136. doi: 10.1021/acscentsci.9b00397. PMC 6661864. PMID 31404244.
- ^ a b Schwizer F, Okamoto Y, Heinisch T, Gu Y, Pellizzoni MM, Lebrun V, et al. (January 2018). "Artificial Metalloenzymes: Reaction Scope and Optimization Strategies" (PDF). Chemical Reviews. 118 (1): 142–231. doi: 10.1021/acs.chemrev.7b00014. PMID 28714313.
- ^ Akabori S, Sakurai S, Izumi Y, Fujii Y (August 1956). "An Asymmetric Catalyst". Nature. 178 (4528): 323–324. Bibcode: 1956Natur.178..323A. doi: 10.1038/178323b0. ISSN 0028-0836. PMID 13358737. S2CID 4221816.
- ^ Wilson ME, Whitesides GM (January 1978). "Conversion of a protein to a homogeneous asymmetric hydrogenation catalyst by site-specific modification with a diphosphinerhodium(I) moiety". Journal of the American Chemical Society. 100 (1): 306–307. doi: 10.1021/ja00469a064. ISSN 0002-7863.
- ^ Yamamura K, Kaiser ET (1976-01-01). "Studies on the oxidase activity of copper(II) carboxypeptidase A". Journal of the Chemical Society, Chemical Communications (20): 830–831. doi: 10.1039/C39760000830. ISSN 0022-4936.
- ^ Davies RR, Distefano MD (1997-12-01). "A Semisynthetic Metalloenzyme Based on a Protein Cavity That Catalyzes the Enantioselective Hydrolysis of Ester and Amide Substrates". Journal of the American Chemical Society. 119 (48): 11643–11652. doi: 10.1021/ja970820k. ISSN 0002-7863.
- ^ Onoda A, Kihara Y, Fukumoto K, Sano Y, Hayashi T (August 2014). "Photoinduced Hydrogen Evolution Catalyzed by a Synthetic Diiron Dithiolate Complex Embedded within a Protein Matrix". ACS Catalysis. 4 (8): 2645–2648. doi: 10.1021/cs500392e.
- ^ Davies RR, Distefano A (December 1997). "A Semisynthetic Metalloenzyme Based on a Protein Cavity That Catalyzes the Enantioselective Hydrolysis of Ester and Amide Substrates". Journal of the American Chemical Society. 119 (48): 11643–11652. doi: 10.1021/ja970820k. ISSN 0002-7863.
- ^ Platis IE, Ermácora MR, Fox RO (November 1993). "Oxidative polypeptide cleavage mediated by EDTA-Fe covalently linked to cysteine residues". Biochemistry. 32 (47): 12761–7. doi: 10.1021/bi00210a027. PMID 8251497.
- ^ Yang H, Srivastava P, Zhang C, Lewis JC (January 2014). "A general method for artificial metalloenzyme formation through strain-promoted azide-alkyne cycloaddition". ChemBioChem. 15 (2): 223–7. doi: 10.1002/cbic.201300661. PMC 3996923. PMID 24376040.
- ^ Kruithof CA, Casado MA, Guillena G, Egmond MR, van der Kerk-van Hoof A, Heck AJ, et al. (November 2005). "Lipase active-site-directed anchoring of organometallics: metallopincer/protein hybrids". Chemistry: A European Journal. 11 (23): 6869–77. doi: 10.1002/chem.200500671. PMID 16224766.
- ^ Creus M, Pordea A, Rossel T, Sardo A, Letondor C, Ivanova A, et al. (2008). "X-ray structure and designed evolution of an artificial transfer hydrogenase". Angewandte Chemie. 47 (8): 1400–4. doi: 10.1002/anie.200704865. PMID 18176932.
- ^ Creus M, Pordea A, Rossel T, Sardo A, Letondor C, Ivanova A, et al. (2008-02-08). "X-Ray Structure and Designed Evolution of an Artificial Transfer Hydrogenase". Angewandte Chemie International Edition. 47 (8): 1400–1404. doi: 10.1002/anie.200704865. ISSN 1433-7851. PMID 18176932.
- ^ Iverson BL, Lerner RA (March 1989). "Sequence-specific peptide cleavage catalyzed by an antibody". Science. 243 (4895): 1184–8. Bibcode: 1989Sci...243.1184I. doi: 10.1126/science.2922606. PMID 2922606.
- ^ Roelfes G (2019-02-22). "LmrR: A Privileged Scaffold for Artificial Metalloenzymes". Accounts of Chemical Research. 52 (3): 545–556. doi: 10.1021/acs.accounts.9b00004. ISSN 0001-4842. PMC 6427492. PMID 30794372.
- ^ Villarino L, Chordia S, Alonso-Cotchico L, Reddem E, Zhou Z, Thunnissen AM, et al. (2020-09-18). "Cofactor Binding Dynamics Influence the Catalytic Activity and Selectivity of an Artificial Metalloenzyme". ACS Catalysis. 10 (20): 11783–11790. doi: 10.1021/acscatal.0c01619. ISSN 2155-5435. PMC 7574625. PMID 33101759.
- ^ Zhou Z, Roelfes G (2021-07-13). "Synergistic Catalysis of Tandem Michael Addition/Enantioselective Protonation Reactions by an Artificial Enzyme". ACS Catalysis. 11 (15): 9366–9369. doi: 10.1021/acscatal.1c02298. ISSN 2155-5435. PMC 8353628. PMID 34386272.
- ^ "Graphical Abstract: Angew. Chem. Int. Ed. 17/2018". Angewandte Chemie International Edition. 57 (17): 4435–4453. 2018-04-16. doi: 10.1002/anie.201881711. ISSN 1433-7851.
- ^ Kandemir B, Chakraborty S, Guo Y, Bren KL (January 2016). "Semisynthetic and Biomolecular Hydrogen Evolution Catalysts". Inorganic Chemistry. 55 (2): 467–77. doi: 10.1021/acs.inorgchem.5b02054. PMID 26671416.
- ^ a b Drienovská I, Roelfes G (2020-01-06). "Expanding the enzyme universe with genetically encoded unnatural amino acids". Nature Catalysis. 3 (3): 193–202. doi: 10.1038/s41929-019-0410-8. ISSN 2520-1158.
- ^ Drienovská I, Rioz-Martínez A, Draksharapu A, Roelfes G (January 2015). "Novel artificial metalloenzymes by in vivo incorporation of metal-binding unnatural amino acids". Chemical Science. 6 (1): 770–776. doi: 10.1039/C4SC01525H. PMC 5590542. PMID 28936318.
- ^ Drienovská I, Scheele RA, Gutiérrez de Souza C, Roelfes G (November 2020). "A Hydroxyquinoline-Based Unnatural Amino Acid for the Design of Novel Artificial Metalloenzymes". ChemBioChem. 21 (21): 3077–3081. doi: 10.1002/cbic.202000306. PMC 7689906. PMID 32585070.
- ^ Chen K, Huang X, Zhang SQ, Zhou AZ, Kan SB, Hong X, et al. (March 2019). "c-Catalyzed Lactone-Carbene B-H Insertion". Synlett. 30 (4): 378–382. doi: 10.1055/s-0037-1611662. PMC 6436545. PMID 30930550.
- ^ Brandenberg OF, Chen K, Arnold FH (May 2019). "Directed Evolution of a Cytochrome P450 Carbene Transferase for Selective Functionalization of Cyclic Compounds" (PDF). Journal of the American Chemical Society. 141 (22): 8989–8995. doi: 10.1021/jacs.9b02931. PMID 31070908.
- ^ Cho I, Prier CK, Jia ZJ, Zhang RK, Görbe T, Arnold FH (March 2019). "Enantioselective Aminohydroxylation of Styrenyl Olefins Catalyzed by an Engineered Hemoprotein". Angewandte Chemie. 58 (10): 3138–3142. doi: 10.1002/anie.201812968. PMID 30600873.
- ^ a b Zhou Q, Chin M, Fu Y, Liu P, Yang Y (2021-12-24). "Stereodivergent atom-transfer radical cyclization by engineered cytochromes P450". Science. 374 (6575): 1612–1616. Bibcode: 2021Sci...374.1612Z. doi: 10.1126/science.abk1603. ISSN 0036-8075. PMC 9309897. PMID 34941416.
- ^ Rui J, Zhao Q, Huls AJ, Soler J, Paris JC, Chen Z, et al. (2022-05-20). "Directed evolution of nonheme iron enzymes to access abiological radical-relay C(sp 3 )−H azidation". Science. 376 (6595): 869–874. doi: 10.1126/science.abj2830. ISSN 0036-8075. PMC 9933208. PMID 35587977.
- ^ a b Zhao Q, Chen Z, Soler J, Chen X, Rui J, Ji NT, et al. (2024-03-28). "Engineering non-haem iron enzymes for enantioselective C(sp3)–F bond formation via radical fluorine transfer". Nature Synthesis. doi: 10.1038/s44160-024-00507-7. ISSN 2731-0582.
- ^ a b Yang Y, Zhao LP, Mai BK, Cheng L, Gao F, Zhao Y, et al. (2024-03-29). "Biocatalytic enantioselective C(sp3)–H fluorination enabled by directed evolution of nonheme Fe enzymes". dx.doi.org. doi: 10.26434/chemrxiv-2024-pt58m. Retrieved 2024-06-07.
- ^ Dydio P, Key HM, Nazarenko A, Rha JY, Seyedkazemi V, Clark DS, et al. (October 2016). "An artificial metalloenzyme with the kinetics of native enzymes". Science. 354 (6308): 102–106. Bibcode: 2016Sci...354..102D. doi: 10.1126/science.aah4427. PMID 27846500.
- ^ Yokoi N, Inaba H, Terauchi M, Stieg AZ, Sanghamitra NJ, Koshiyama T, et al. (September 2010). "Construction of robust bio-nanotubes using the controlled self-assembly of component proteins of bacteriophage T4". Small. 6 (17): 1873–9. doi: 10.1002/smll.201000772. PMID 20661999.
- ^ Key HM, Dydio P, Clark DS, Hartwig JF (June 2016). "Abiological catalysis by artificial haem proteins containing noble metals in place of iron". Nature. 534 (7608): 534–7. Bibcode: 2016Natur.534..534K. doi: 10.1038/nature17968. PMID 27296224. S2CID 205249065.
- ^ Podtetenieff J, Taglieber A, Bill E, Reijerse EJ, Reetz MT (July 2010). "An artificial metalloenzyme: creation of a designed copper binding site in a thermostable protein". Angewandte Chemie. 49 (30): 5151–5. doi: 10.1002/anie.201002106. PMID 20572232.
- ^ Drienovská I, Rioz-Martínez A, Draksharapu A, Roelfes G (January 2015). "in vivo incorporation of metal-binding unnatural amino acids". Chemical Science. 6 (1): 770–776. doi: 10.1039/C4SC01525H. PMC 5590542. PMID 28936318.
- ^ Arnold FH (April 2018). "Directed Evolution: Bringing New Chemistry to Life". Angewandte Chemie. 57 (16): 4143–4148. doi: 10.1002/anie.201708408. PMC 5901037. PMID 29064156.
- ^ Renggli K, Nussbaumer MG, Urbani R, Pfohl T, Bruns N (January 2014). "A chaperonin as protein nanoreactor for atom-transfer radical polymerization". Angewandte Chemie. 53 (5): 1443–7. doi: 10.1002/anie.201306798. PMID 24459061.